Improving PROTAC properties via single-point changes to linkers
We explore how computational methods can be applied to proteolysis targeting chimera (PROTAC) design, to effectively tackle some of the ...
News
We are delighted to announce the release of our latest suite of KNIME™ nodes. This release brings Cresset’s new Activity Miner module into the KNIME environment together with a number of improvements.
Activity Miner is designed to find activity cliffs in your Structure Activity Relationships. It uses the disparity between two molecules (difference in activity divided by distance between the molecules) to highlight where small changes in structure result in disproportionately large changes in activity. Uniquely, Activity Miner operates using both 3D and 2D similarity metrics in assessing the change in a structure enabling a full understanding of the reasons for the change in activity.
The Activity Miner KNIME node is part of Cresset’s Forge nodes which wrap the corresponding functionality in the Forge command lines. It operates in three modes, generating either a similarity, distance or disparity matrix for all molecules. When loaded with 2D molecules, the node uses a 2D similarity value to output the matrices. However, loading pre-aligned 3D molecules, such as those from the Forge_align node enables the use of 3D similarity in the Activity Miner module to output matrices based on the biological context in which the change in activity is observed.
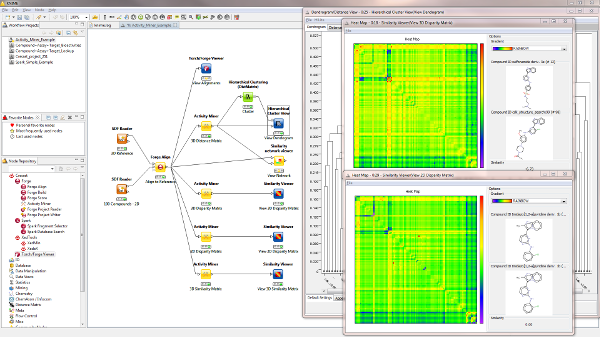
Use the new Activity Miner node in KNIME to extend your existing similarity based workflows to include Cresset’s 3D similarity, bringing the biological context of your molecules into your cheminformatics based decisions.