Before working at Cresset I used Cresset software in the highly dynamic pharmaceutical ‘fee for service’ environment at BioFocus where I supported medicinal chemistry and a diverse range of programs. In this post I describe what it is like to use Cresset software on service projects.
Why people choose Cresset
Cresset has a combination of very clever science and ‘just as clever’ software design that, in my opinion, is a winning combination. As a Cresset employee, I’m proud to say that I was an enthusiastic user of Cresset software before I joined the fold.
Cresset is well known as a provider of cutting edge software for molecular modeling. Our unique science for generating molecular electrostatic fields is deployed in our range of software Blaze, Forge, Spark and Torch. Blaze, our powerful virtual screening platform, has been adopted as a standard virtual screening technique by many of the pharmaceutical giants. Our other software suites have been carefully designed to use Cresset scientific methods to great effect for numerous drug discovery applications. These include: alignment hypotheses generation, scaffold hopping, fragment replacement, iterative design, 3D-QSAR, data visualisation, multi parameter optimization and so on. Many of these tasks are more often associated with computational chemistry and confined to a small enclave of the drug discovery community.
The Cresset philosophy on small molecule discovery and software has always been ‘inclusive’, appealing to both expert computational chemists and experienced medicinal and synthetic chemists who are drawn to our software because it is intuitive and easy to use as well as extremely powerful.
Services as well as software for discovery research
What is less well known about Cresset is that for as long as we have been creating excellent software, we have been actively engaged in providing modeling support for a wide variety of molecular discovery programs. The term ’molecular discovery’ is deliberately vague; this is because the breadth of this activity covers pharmaceutical research, agrochemicals, flavors, fragrances and chemicals. Any endeavour involving molecular recognition comes under the umbrella.
Looking back over the last few years it would not be surprising to note that virtual screening is amongst the most popular tasks we are asked to perform. Scaffold hopping and fragment replacement are a close second and latterly SAR analyses such as 3D-QSAR have also become increasingly popular as these functionalities have been made available. Partly this has been driven by an increased awareness of the ability of Cresset Discovery Services to deliver these analyses as our commercial software offering has strengthened and grown.
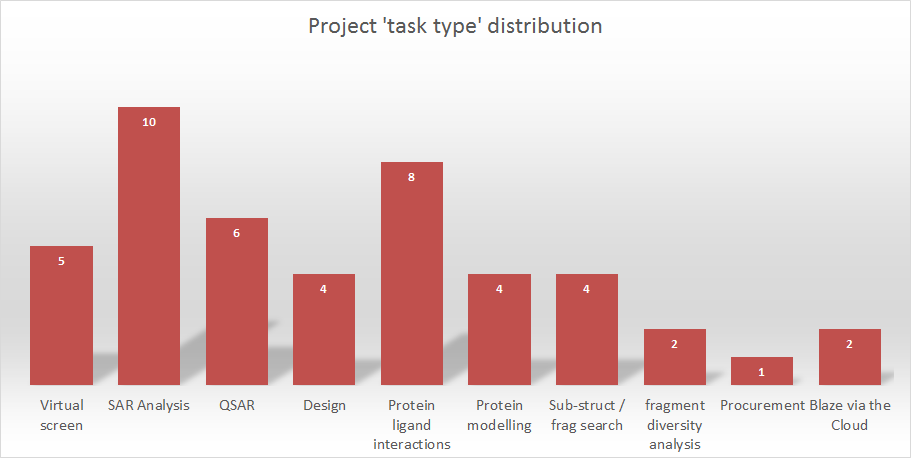
Figure 1: Task distribution for service projects
Conversely, and possibly a little more surprising is that some protein-centric rather than ligand-centric analyses form an increasing proportion of our services business. Protein-ligand interactions, homology modeling and protein-ligand minimization are becoming frequent tasks. The simple fact is that the Cresset XED force field and electrostatics offer a different view of proteins, in the same way that our ligand centric analyses offer important insights that can’t be found using other methods.
Currently the Cresset Discovery Services team uses proprietary in-house software for protein modeling. Some of these functionalities and additional new science will be released in due course. In the meantime we continue to offer these activities as a service only.
Molecular recognition detective work
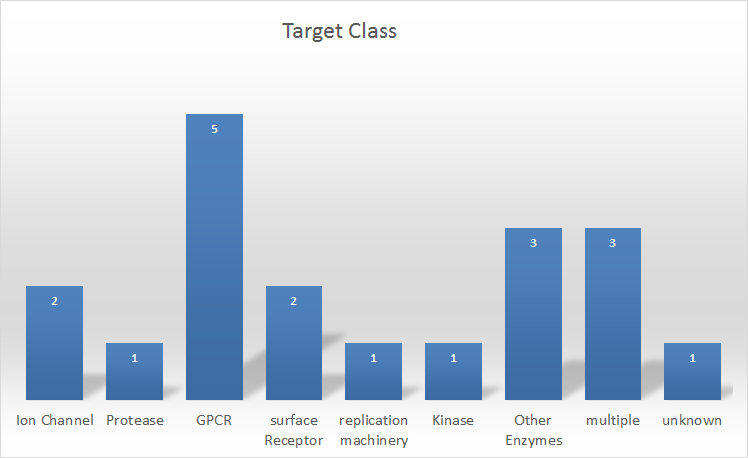
Figure 2: Protein target class distribution for service projects
In terms of the type of protein-ligand systems we are most often engaged with, historically we frequently worked in areas for which structural data was limited such as GPCRs and ion channels. This trend continues despite the increasing structural data available for both areas. We are experiencing something close to a renaissance in GPCR research and ion channel research through the development of new X-ray crystallography and cryo-electron microscopy techniques. See Andy Vinter’s work on
Applying the XED molecular mechanics force field to the binding mechanism of GPCRs.
The calcium channel Cav 1.1 is one such new structure; and for anyone who has worked on ion channels it’s a fascinating structure to now have. Both the sodium Nav and calcium Cav channels are distinct from potassium Kv channels in that they are heterotetrameric, rather than homotetrameric. This raises the possibility that the ordering of sub-units of the tetramer can either be clockwise or anti-clockwise. This is a super puzzle and very much the type of molecular recognition detective work that typifies ‘life at the coal face’ for us.
Through careful consideration of ligand binding hypotheses in conjunction with mutagenesis data it is possible to deduce that the clockwise arrangement for both Nav and Cav channels is the most consistent with the data – which is now proven to be the case.
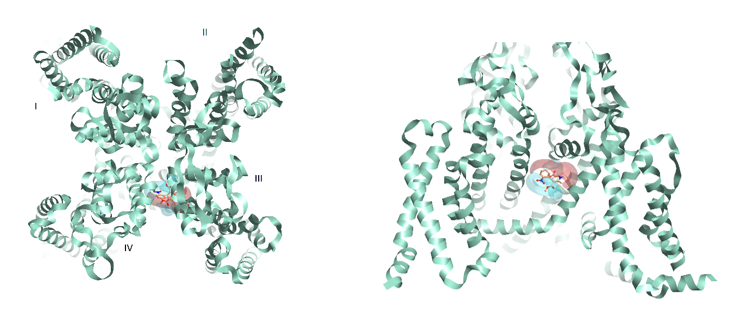
Figure 3: Rabbit Cav1.1 (PDB:3JBR) with Nifedipine and corresponding positive and negative electrostatics. From extracellular view (left) from the side (right) flipped 90 degrees.
Figure 3 shows a binding hypothesis for the dihydropyridine cardiovascular drug Nifedipine in the Cav1.1 binding site groove between domain III and IV. Positive and negative electrostatics are consistent with residues in the local environment and the geometry of the channel.
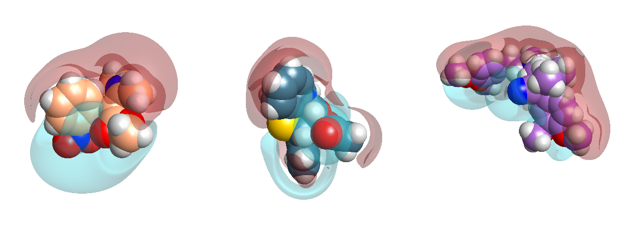
Figure 4: Close-up electrostatic fields of Nifedipine, Diltiazem and Verapamil.
The overall dipole is ‘with’ the ion channel P-loop helix. Interestingly this dipole is exhibited by two other classic calcium channel drugs: verapamil and diltiazem, which bind in the adjacent I and IV domain groove (Figure 4).
The variety of work we engage in means that there are always new challenges. The most exciting times for me are when we apply Cresset methods to new areas and actually manage to elucidate or predict interactions and mechanisms that are later tested and confirmed by experimentation.
Find out more
Whether you need a fully resourced partner for a long-term collaboration, or specific expertise for a focused short-term project, contact us for a discussion about how we can advance your research.



