Improving PROTAC properties via single-point changes to linkers
We explore how computational methods can be applied to proteolysis targeting chimera (PROTAC) design, to effectively tackle some of the ...
News
Flare V2 is in the final rounds of testing, which means the release announcement is imminent. Ahead of the user group meeting, where we will be presenting this major advancement, this post takes a sneak peek at some of the new features in this version.
Completely rewritten surface generation code results in faster and better surfaces with quality options built in to the surface creation dialog. This is combined with new coloring options for new surfaces to give you more insights into your proteins and ligands.
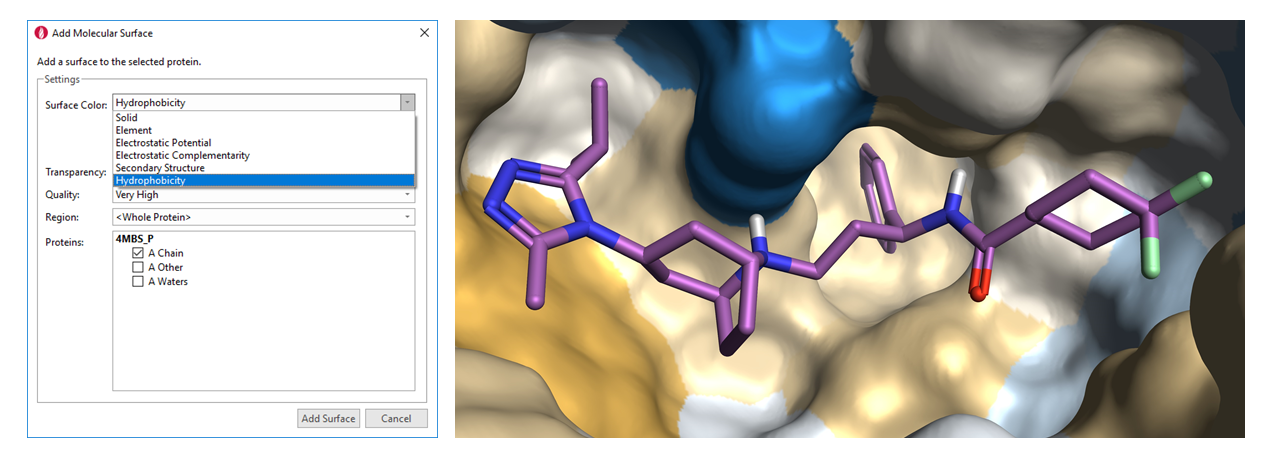
Figure 1: (a) New surface coloring options in Flare V2, and (b) PDB code 4MBS with a hydrophobic surface colored yellow (hydrophobic) to blue (hydrophilic).
Making pictures is key to communicating your insights on protein-ligand binding. Flare V2 has major improvements to the Z-clipping to enable you to get the view that you want. In addition, to apply a specific clipping plane to an individual surface, you now have the option to exclude ligands from the clip altogether. This option makes a significant impact on pictures of binding sites that are completely buried.
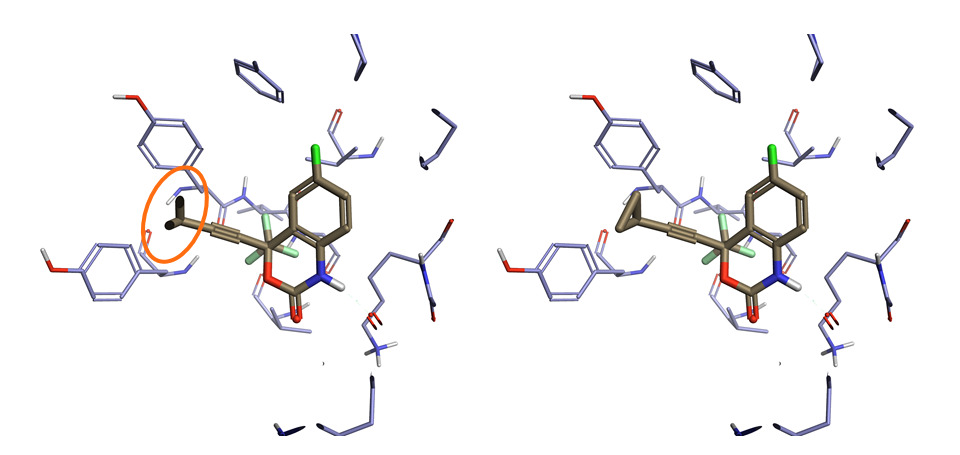
Figure 3: Flare V2 gives the option to clip individual surfaces independently of other objects. Here a clipping plane is added only to the electrostatic surface enabling the visualization of protein residues that are above the ligand in combination with a surface.
The new protein surfaces are complemented by new options for ligand surfaces, the new storyboard panel to capture and replay key 3D insights and many new features for ligands. Taken together with the Python API this release of Flare is a major advancement in this innovative new application for structure-based design.
Register for the up-coming user group meeting to find out more about Flare V2, network with existing users and receive free training at one of the hands-on workshops.