Improving PROTAC properties via single-point changes to linkers
We explore how computational methods can be applied to proteolysis targeting chimera (PROTAC) design, to effectively tackle some of the ...
News
We believe that the lively academic environment is an amazing source of new scientific ideas, algorithms and computational methods. Flare for Academics is a free* licensing option of Flare, our structure-based design software, which has specifically been designed for academic users.
Flare for Academics is a user-friendly environment where academic users can easily develop and test their ideas and methods, or plug-in the most interesting open-source algorithms. It extends on the functionality of Flare Viewer to provide an excellent platform for drug discovery, with a focus on ligand design and electrostatics.
The Flare Python API gives academic researchers the opportunity to make their science more accessible through integration into a user-friendly environment.
You will benefit from a robust, commercial standard SBDD environment that enables focus on science by utilizing methods such as protein preparation, protein minimization and multi-core docking. Access is also given to the RDKit cheminformatics toolkit, NumPy, SciPy, and Matplotlib, which are all integral to Flare. Beyond these, virtually any other Python module can be pip-installed making Flare infinitely extendable. An ever-growing collection of featured python extensions that enhance the existing Flare functionality are also provided, these include: plotting, protein mutation, and custom workflows (see also the new Jupyter Notebook integration).
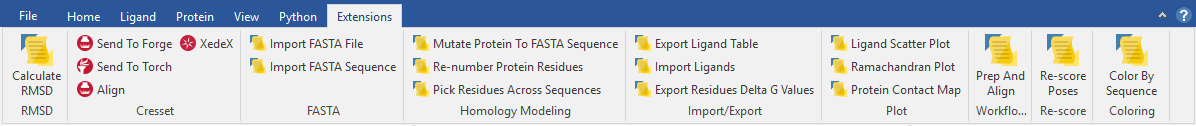
The Flare Python API not only provides an environment to develop your own algorithms but also a way to deploy them across a wider user base. The API provides access to all elements of the Flare interface through addition of user-defined controls and context menus.
For example, you may add custom controls into an existing Flare ribbon, or create a new Flare ribbon for Python scripts you frequently use. Custom-created controls in Flare can be created as small or large buttons, spin boxes, custom sliders, or complex dialogues with signals and call-back functions (Figure 2).
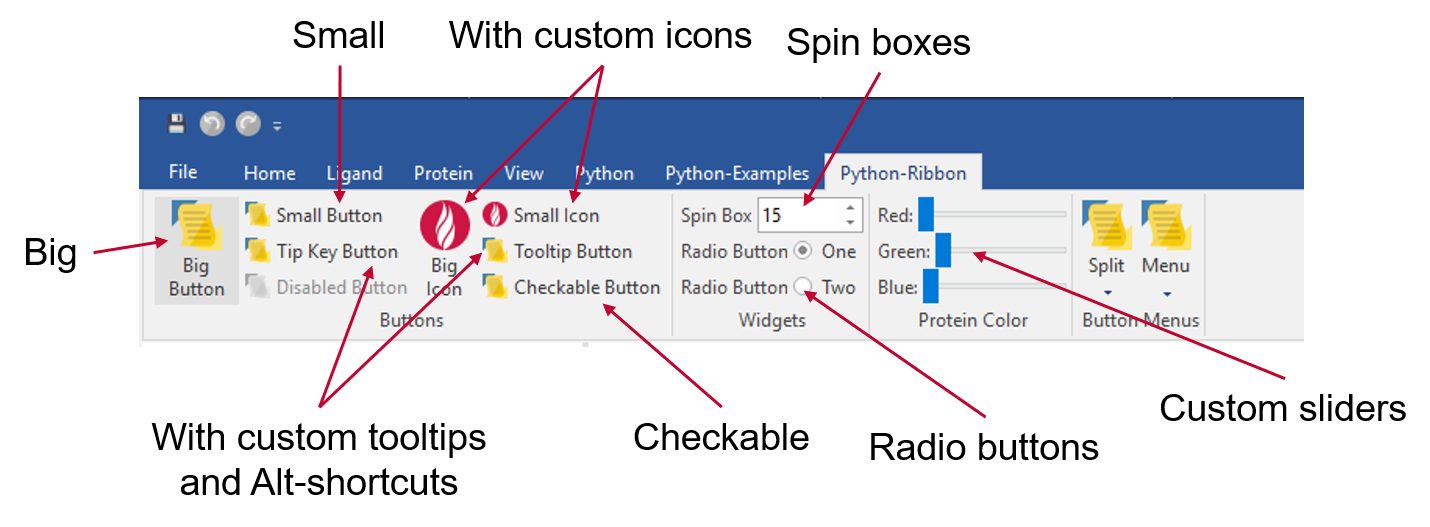
Whenever you need to carry out a completely automated task, for example the overnight preparation of a panel of proteins followed by docking of several ligand series, the most convenient option is to write a Python script that runs outside the Flare GUI. It can then be distributed on a cluster via a queueing system for maximum performance. The pyflare binary is a Python interpreter giving you access to Flare functions using either custom developed or Cresset released scripts.
The native GUI of Flare embeds the Python Console and Python Interpreter widgets. The Python Console is the simplest option to run one-line commands. With the Python Interpreter you can handle slightly more complex scripts: for example, you can load a script, interactively edit it inside Flare and then save your modifications. Both the Python Console and the Python Interpreter have a multi-tab interface that makes it possible to work on multiple Python snippets at the same time.
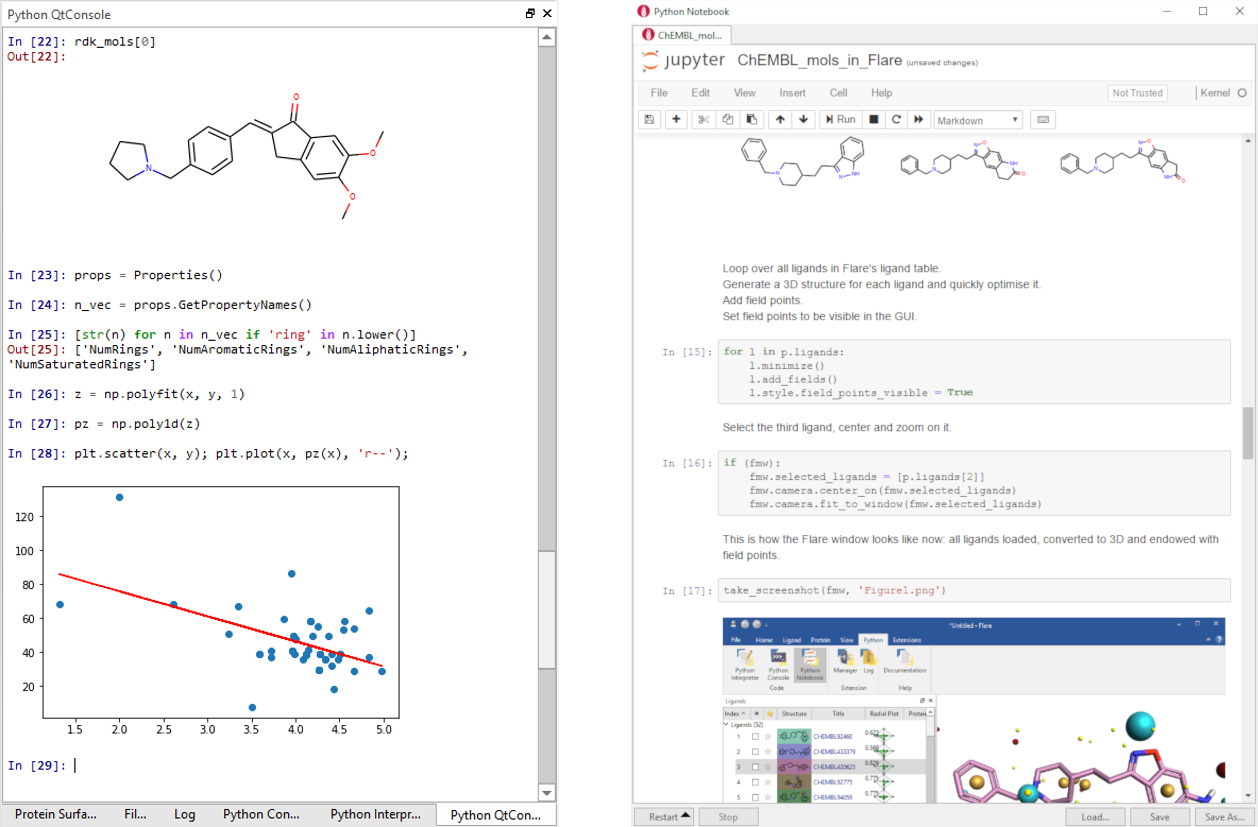
Python enthusiasts can easily upgrade Flare with the Jupyter QtConsole for access to all the Jupyter features, e.g.: TAB completion, auto-indentation, syntax highlighting, context help, inline graphics, and more. Using this widget, you can type Python commands, examine molecules and draw plots, all in the same window.
The Flare Python Notebook is an instance of the Jupyter Notebook embedded into the Flare GUI. It has direct access to the Flare GUI objects and methods, offers an even richer interface and enables editing and running individual code cells.
Flare for Academics is not just a viewer, but a complete, user friendly platform for iterative molecule design in drug discovery.
Multiple protein structures can be easily imported in the Flare project and displayed in the same frame of reference using the sequence alignment and superimposition functions.
Flare’s protein preparation will enable you to optimize your protein-ligand structures by adding hydrogen atoms, optimizing hydrogen bonds, removing atomic clashes and assigning optimal protonation states. Further optimization of the protein active site can be achieved by protein minimization based on the XED force field, and by manually flipping flexible residues or changing tautomeric and charge states for relevant residues.
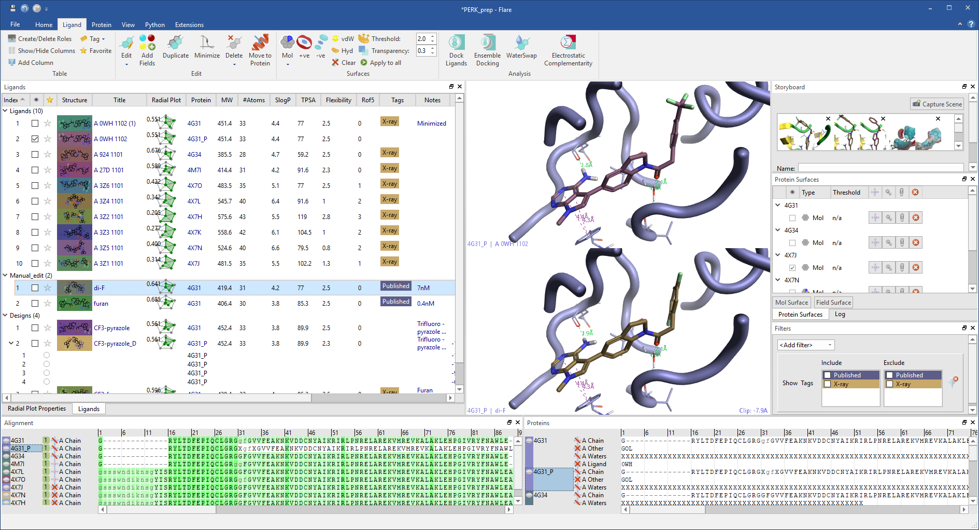
Smart visualization of protein-ligand complexes in grid mode facilitates the comparison between ligand or proteins. The display of a variety of non-bonded ligand-protein interactions makes it easy to understand the different binding modes for your ligands.
The ligand-centric structure of Flare includes a dedicated ligand table and interactive menu giving easy access to all ligand actions, such as sorting on any column, control visibility, tagging and filtering on structure, tags and numerical and text columns, grouping of ligands in custom-created roles. In the ligand table, each molecule is associated to calculated physico-chemical properties, a radial plot and a multi-parametric score to help you design and select the ligands that best match the ideal project profile. Ligand electrostatic interaction potentials calculated with the XED force field can be visualized in the 3D window and in the molecule editor, and used to inform ligand design.
Multi-core docking experiments can be run to predict the 3D structure of flexible ligands in the active site of your protein. Docking in Flare uses Lead Finder™ to provide excellent pose prediction and detailed feedback on new molecule designs.
See the features of Flare for Academics, and apply for your 1 year license.
* In most countries; contact us to see if you are eligible for a free license.