By the time we are 65, one in ten of us will have Alzheimer’s disease. This rises to fifty percent by the age of 85 and it is predicted that by 2050 there will be 130 million people worldwide living with Alzheimer’s disease. Care for Alzheimer’s patients tends to be complex, intensive and prolonged which places an ever-increasing burden on health services, families and support staff. Currently there are only four FDA approved drugs for the alleviation of the symptoms of Alzheimer’s disease but none that actually treat the disease.
Alzheimer’s disease1 is characterized by the formation of plaques and tangles in the brain leading to neuronal cell damage and death. These are formed by Aβ42 proteins and hyperphosphorylated Tau proteins respectively2,3. Drug discovery for Alzheimer’s disease has focused on preventing the formation of these plaques and tangles but has been hampered by the difficulty in identifying potential drug targets and by the complexity of running clinical trials. Several groups have begun working on identifying alternative potential targets which could be addressed to prevent or remove the observed Alzheimer’s related plaques.
Computational chemistry is a valuable resource in helping to characterize and understand new protein targets. Cresset Discovery Services uses a wide and continually growing set of applications in both the ligand and structure-based fields to give your research groups unrivaled insight into new targets.
How to model a disordered protein
The Tau protein is emerging as both a biomarker and potentially therapeutic target for Alzheimer’s disease as well as other neuro degenerative diseases. These proteins are primarily found in the neurons of the central nervous system where their function is to stabilize microtubules. Hyperphosphorylation of the Tau protein can result in self-assembly of filaments which are involved in the development of Alzheimer’s disease. Compounds which alter the stability and ability of Tau to self-assemble or prevent the hyperphosphorylation could therefore have a role in treating Alzheimer’s disease.
Tau proteins are classed as disordered proteins, so their 3D structure is strongly determined by their binding partner. If this binding partner is known it could be used to postulate a bound Tau structure, but it is more likely that this target would be treated in a ligand-based manner driven by developing QSAR models around a set of active compounds. Using the QSAR model building capabilities of Forge™ Cresset Discovery Services can generate binding mode hypotheses and use this knowledge with the project team’s combined experience to propel the project forward.
Lipoxygenases, caspases and sirtuins
Crystal structures are available for Lipoxygenases, Caspases and Sirtuins. They can therefore be modeled using ligand-based methods to develop a homology model in Forge, or by using a structure-based approach in Flare™, with the identification of potential active or allosteric sites followed by docking in Lead Finder.
Lipoxygenases
Lipoxygenases are a group of non-heme iron containing dioxygenases that catalyze the addition of oxygen to a range of polyunsaturated fatty acids (Figure 1). These compounds act as signalling molecules for a wide variety of cell functions including neural inflammation. Family members 12LO and 15LO are linked to fatty acids oxidation signalling pathways in neurodegeneration. Increased levels have been observed in areas susceptible to Alzheimer’s disease. Lipoxygenases are up-steam of both Aβ42 and Tau hyperphosphorylation so could have beneficial effects against both Aβ42 plaques and Tau hyperphosphorylation tangles, making them an exciting target.
![Ribbon for the 2p0m crystal structure 2p0m a revised structure of rabbit reticulocyte 15S-lipoxygenase [4], created using Flare™. ribbon for the 2p0m crystal structure 2p0m](/media/cache/f9/47/f94732279f1dc599f1face9140bed6e5.png)
Figure 1: Ribbon for the 2p0m crystal structure 2p0m a revised structure of rabbit reticulocyte 15S-lipoxygenase 4, created using Flare™.
Caspases
Caspases are a group of cystenyl proteases that are part of the apoptotic pathway. Several studies have suggested a role for caspases in the Aβ and Tau hyperphosphorylation neurodegenerative pathways. Caspase-1 has been shown in mice to reduce Aβ levels and reduce brain inflammation in the J20 mouse mode of Alzheimer’s disease (Figure 2) 5.
![Ribbon for the 1ice Crystal structure of the human caspase -1.[6], created using Flare. Ribbon for the 1ice Crystal structure of the human caspase](/media/cache/a8/e2/a8e2daee48aaef530643eb857b17d391.png)
Figure 2: Ribbon for the 1ice Crystal structure of the human caspase -16, created using Flare.
Sirtuins
Sirtuins are from the family of nicotinamide dependent protein deacetylases that are involved with a large range of cellular processes (Figure 3). There is a growing amount of evidence of the importance of SIRT1, and the full family of sirtuin proteins in Alzheimer’s disease prevention and diagnosis. Currently there is only one crystal structure of SIRT2 from the sirtuins family which, despite the low sequence homology across the family, could be used to generate homology models of the remaining family members.
![Ribbon for the crystal structure 5yql human Sirtuin-2 the only crystal structure available for the sirtuin family with the selective inhibitor A2I [7], created using Flare. Ribbon for the crystal structure 5yql human Sirtuin-2](/media/cache/41/5e/415e33518382464326834114041cd96d.png)
Figure 3: Ribbon for the crystal structure 5yql human Sirtuin-2 the only crystal structure available for the sirtuin family with the selective inhibitor A2I7, created using Flare.
The secretase pathway: Removing unwanted side effects
Three secretase proteins are involved across two distinct biological pathways. Both process the amyloid precursor protein: the α-γ-secretase pathway which is non-amyloidogenic, and the β-γ Secretase pathway which is amyloidogenic (Figure 4).
The secretase systems are potentially open to both structure or ligand-based approaches; however, these present the challenge of removing NOTCH-1 and other off-target activities which are responsible for the side-effects that ended the clinical trials.
Activity Atlas™ can be used to develop multiple QSAR models of a system. This can help to identify chemical functionality that could remove the unwanted activities whilst protecting the favorable activity and increasing the therapeutic window.
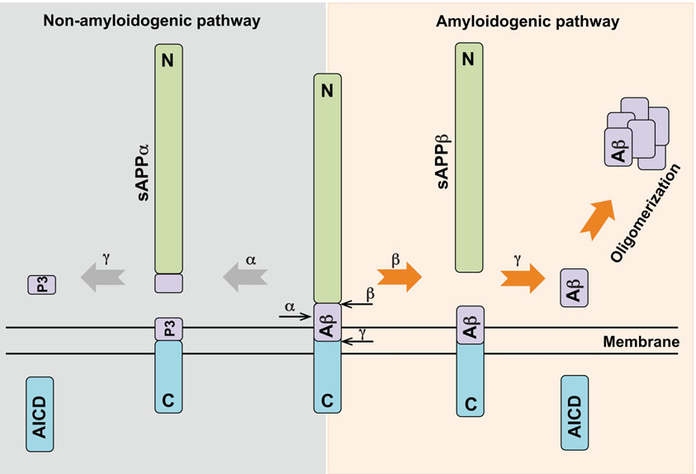
Figure 4: The two secretase pathways: α-γ secretase and β-γ secretase8.
α-Secretase
The amyloid precursor protein is cleaved by α-secretase to generate a shorter P3 system. This is non-amyloidogenic so if this pathway could be promoted it offers a mechanism for lowing the production of αβ429.
β-Secretase
The β-site amyloid precursor protein-cleaving enzyme 1 (BASE1 or β-secretase) is a member of the pepsin-like family of aspartyl proteases and cleavages amyloid precursor protein. If there is less amyloid precursor protein cleaved then less Aβ42 can be produced. Several drugs have been directed towards this goal and there has been some evidence that inhibition does work in reducing Aβ42 levels. However, there have also been several side effects which have resulted in the termination of clinical trials for several compounds. This is because BASE1 also has multi-substrate specificity including the NOTCH 1 pathway. Partial inhibition or selective inhibition of the amyloid precursor protein binding site may be possible to reduce the number of side effects, but this remains unproved.
γ-Secretase
γ-Secretase is a complex composed of 4 subunits which act as regulators: nicastrin, presenilin-1 (PSEN1), anterior pharynx-defective-1 (APH-1), presenilin enhancers 2 and CD147. A γ-secretase inhibitor DAPT has been shown to be effective in reducing levels of Aβ present in the brain10. γ -secretase is also part of the NOTCH 1 pathway and gaining selectivity for only the Aβ cleavage has hampered this avenue of research. Directly inhibiting γ-secretase seems to generate a host of unwanted side effects, but modulating the regulatory mechanisms via CD147, γ-secretase activating protein, etc. may present a more tolerated method of controlling γ-secretase and reduce the production of Aβ units.
Find out more about modeling new protein targets
Cresset Discovery Services has a wealth of experience across multiple protein families. If you have found a novel target, contact us for a free confidential discussion to see how to progress your project.
- A. Alzheimer, Uber eine eigenartige Erkrankung der Hirnrinde, Zentralbl. Nervenh. Psych. 18 (1907) 177–179.
- Glenner GG and Wong CW (1984) Alzheimer’s disease and Down’s syndrome: Sharing of a unique cerebrovascular amyloid fibril protein. Biochem Biophys Res Commun 122, 1131-1135.
- Masters CL, Simms G, Weinman NA, Multhaup G, McDonald BL, Beyreuther K (1985) Amyloid plaque core protein in Alzheimer disease and Down syndrome. Proc Natl Acad Sci U S A 82, 4245-4249.
- Choi, J., Chon, J.K., Kim, S., Shin, W., Conformational flexibility in mammalian 15S-lipoxygenase: Reinterpretation of the crystallographic data., (2008) Proteins 70: 1023-1032.
- Joseph Flores, Anastasia Noël, Bénédicte Foveau, Jeffrey Lynham, Clotilde Lecrux & Andréa C. LeBlanc., Caspase-1 inhibition alleviates cognitive impairment and neuropathology in an Alzheimer’s disease mouse model. Nature Communications volume 9, Article number: 3916 (2018).
- Wilson, K.P., Black, J.A., Thomson, J.A., Kim, E.E., Griffith, J.P., Navia, M.A., Murcko, M.A., Chambers, S.P., Aldape, R.A., Raybuck, S.A., Structure and mechanism of interleukin-1 beta converting enzyme., (1994) Nature 370: 270-275.
- Yang, L.L., Wang, H.L., Zhong, L., Yuan, C., Liu, S.Y., Yu, Z.J., Liu, S., Yan, Y.H., Wu, C., Wang, Y., Wang, Z., Yu, Y., Chen, Q., Li, G.B., X-ray crystal structure guided discovery of new selective, substrate-mimicking sirtuin 2 inhibitors that exhibit activities against non-small cell lung cancer cells. (2018) Eur J Med Chem 155: 806-823.
- Maya Nicolas, Bassem A. Hassan., Amyloid precursor protein and neural development, Development 2014 141: 2543-2548.
- F. Mangialasche, A. Solomon, B. Winblad, P. Mecocci, M. Kivipelto, Alzheimer’s disease: clinical trials and drug development, Lancet Neurol. 9(2010) 702–716.
- M.A. Brodney, D.D. Auperin, S.L. Becker, B.S. Bronk, T.M. Brown, K.J. Coffman,et al., Diamide amino-imidazoles: a novel series of gamma-secretase inhibitors for the treatment of Alzheimer’s disease, Bioorg. Med. Chem. Lett.21 (2011) 2631–2636.
![Ribbon for the 2p0m crystal structure 2p0m a revised structure of rabbit reticulocyte 15S-lipoxygenase [4], created using Flare™. ribbon for the 2p0m crystal structure 2p0m](/media/cache/f9/47/f94732279f1dc599f1face9140bed6e5.png)
![Ribbon for the 1ice Crystal structure of the human caspase -1.[6], created using Flare. Ribbon for the 1ice Crystal structure of the human caspase](/media/cache/a8/e2/a8e2daee48aaef530643eb857b17d391.png)
![Ribbon for the crystal structure 5yql human Sirtuin-2 the only crystal structure available for the sirtuin family with the selective inhibitor A2I [7], created using Flare. Ribbon for the crystal structure 5yql human Sirtuin-2](/media/cache/41/5e/415e33518382464326834114041cd96d.png)
