Future-proofing Cybersecurity in Drug Discovery
The pharmaceutical and biotech sectors suffer more data security breaches than any other industry, with 53% resulting from malicious activity. To protect against potential ...
News
This Fall’s ‘Design A Molecule Competition’ was Cresset’s third and the second to have an Anti-Malarial flavor. Contestants were asked to design a molecule which reconnected the fragments of a Farnesyl Transferase inhibitor from the PDB and to effectively replace the core of the ligand with a new framework. The choice of ligand and target was made with some thought to balance the ease and likely interest in the task with relevance in the field, without getting drawn too deeply into drug the discovery process and its many, many issues.
The very entertaining storm which ensued illustrated perfectly the potentially embroiling diversity of opinion in the drug discovery arena. Despite this and initial reservations about the difficulty of the challenge, Cresset was delighted with the response to the competition. The resultant entry molecules were ranked relative to the original bioactive/bound conformer using Cresset’s ‘3D Molecular Field Similarity’ in FieldAlign and then judged on synthetic tractability and drug-likeness. Novelty was also factored into the ranking by imposing a 2D similarity penalty.
Cresset has no commercial interest in exploiting IP from its competitions and so with consent from contestants we are very happy to disclose the winning structures.
Winner: Congratulations to Peter Ertl of Novartis, the winner of a brand new tablet!
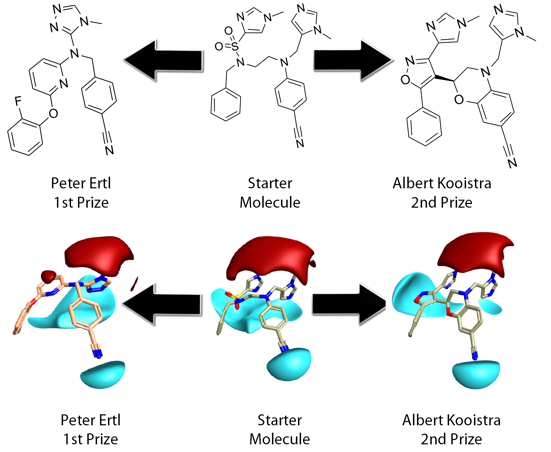
Peter’s design was very elegant, compact, with far less conformational flexibility and high field and shape match, relative to the starting inhibitor. Peter cleverly used the triazole replacement to capture the fields provided by both imidazole moieties in the reference. In the complex these are involved in metal ion and H-bond acceptor interactions from tryptophan/tyrosine. The aryl ether moiety would provide a suitable chemical handle for driving selectivity specifically towards the malarial target. Peter’s design is reasonably drug-like, with a low mwt 400, logP 4.6, TPSA 80 and synthetically tractable.
A very close second place result goes to Albert Kooistra (PhD student in the division of Medicinal Chemistry of VU University Amsterdam). Albert’s design was so near to winning that we have awarded him with a second prize: An Amazon Kindle. Well done!
Albert took a more literal approach, maintaining all the original fragments, to produce high field and shape match, although invoked a higher 2D similarity penalty than Peter. Albert engineered a very nice imidazol-sulphonamide bioisostere which maintains H-bonding interactions to Arg in the complex. These interactions are not as well served as by Peter’s design. Albert’s design is also reasonably drug-like mwt 477, logP 3.9, TPSA 98 but is arguably less synthetically tractable or amenable to further synthetic exploration.
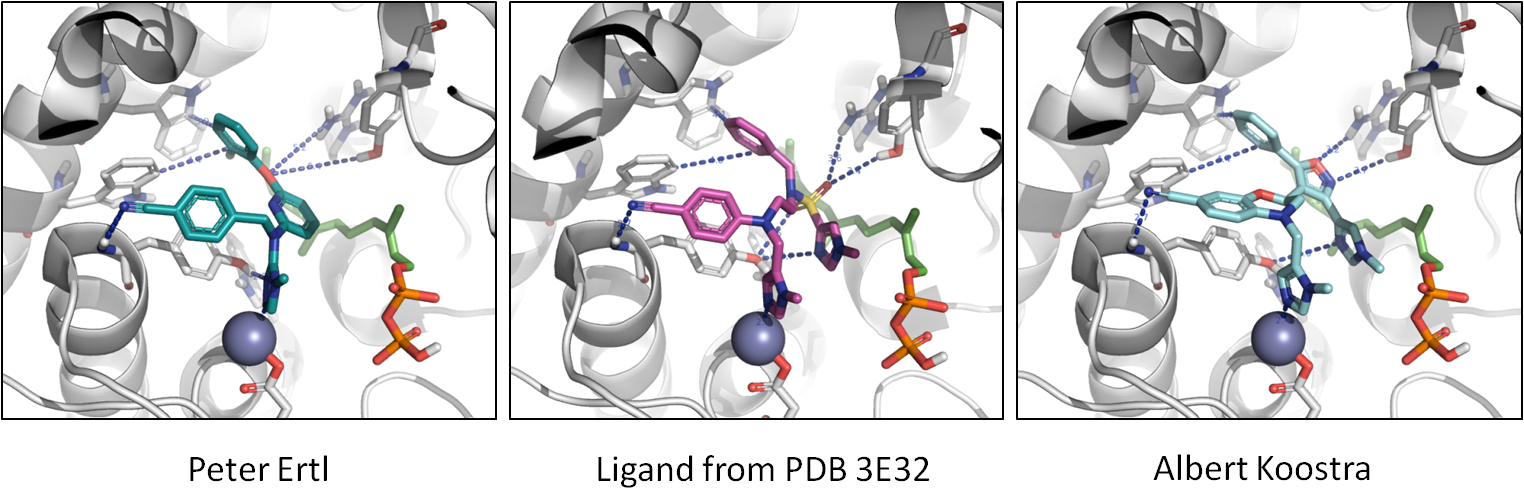
Finally, in the image below, you can see the molecules in the context of protein.
There were several other very nice results worthy of an honourable mention:
Andy Knox (Trinity College Dublin), Brad Teegarden (Consultant), and Sandeep Sundriyal (Texas A&M Health Science Center) all submitted cyano indole and aza-indole starting points, which scored highly and had great tractability. And finally, a very worthy mention for Matt Baumgartner (University of Pittsburgh) who submitted all of his results featuring single step syntheses, showing a commendable appreciation of the issue of cost of goods and synthetic tractability in the Malarial field.
Congratulations to all of our winners and thank you to the rest of our entrants. We will run this competition again, please watch for details.