Improving PROTAC properties via single-point changes to linkers
We explore how computational methods can be applied to proteolysis targeting chimera (PROTAC) design, to effectively tackle some of the ...
News
When I first started using CHARMM over 20 years ago as a PhD student, I regularly came across references to Free Energy Perturbation (FEP); we even discussed it in the group as a possible technique that could be used for an aspect of our work, but the computational resources were lacking at the time to do the technique any justice.
Fast forward 20+ years and we have a different situation. The advent of using Graphic Processor Units (GPUs) to run faster calculations, and access to scalable cloud-based solutions has given more people the computing power required to run these simulations. As well as computational power there have been developments in the force field descriptions and the improvement in conformation sampling techniques which now make FEP calculations a viable technique to use for research in both academia and in industry 1, 2.
FEP is often discussed in terms of two types of calculation:
For the pharmaceutical industry, which is always looking for ways to improve productivity at the hit-to-lead and lead optimization stages, RFEB techniques allow computational and medicinal chemists to shortlist a set of compounds for synthesis and testing. This enables them to potentially reduce the number of compounds that would enter the costly synthesis and testing stages, and so reach the next round of design and development more cost effectively. FEP can, therefore, be a valuable addition to the drug discovery toolbox.
There are inevitably some limitations to the technique. FEP is usually executed using non-physical (‘alchemical’) changes, which gradually morph one ligand into another in a series of steps, often referred to as lambda windows (figure1).
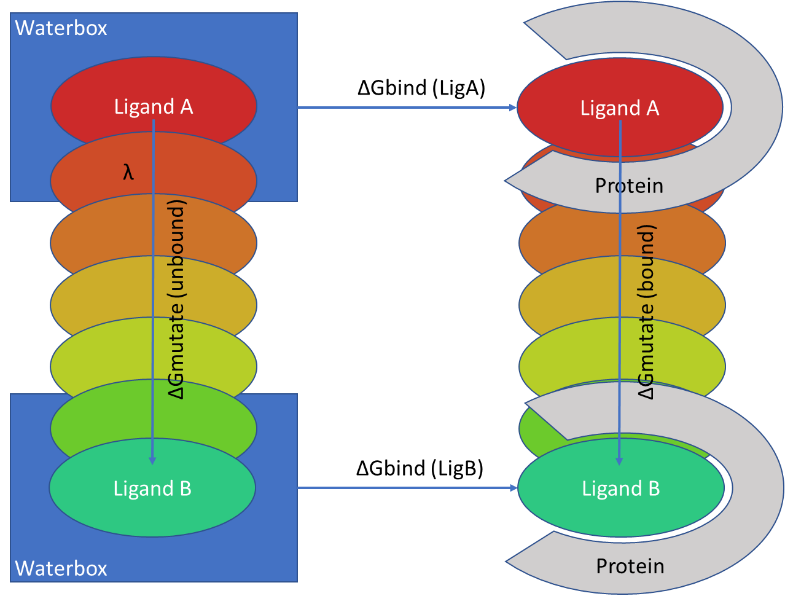
Figure1: Calculating the difference of ΔG binding between Ligand A and Ligand B to a target using alchemical changes.
This requires the FEP process to map atoms between the two ligands, and to ensure that each lambda window adequately overlaps with its neighboring windows, so only limited changes between ligands can be considered with RFEB. In reality, changes between ligands need to be restricted to <10 atoms, so RFEB methods are ideally suited to closely related congenic series, and not scoring a diverse set of compounds coming from a virtual screen.
To run an FEP calculation we need to start with a protein structure, ideally with a bound ligand of interest. The protein structure needs to be prepared so that missing atoms in the side chains are added, as well as missing loops. A protein which is not correctly set-up will probably become unstable in the FEP experiment. Andy Smith, Discovery Services Scientist at Cresset Discovery Services, recently talked about the good practices that should be employed in setting up a model of a biomolecule.
It is important to know the binding mode of the ligand series as this identifies what interactions the ligand series makes with the target protein. FEP simulations tend to have an error of 1kcal mol-1 and so it is important to realize that FEP will not be able to reliably predict the relative ΔG values between ligands with the same order of magnitude of activity, but it would be able to distinguish between micromolar and nanomolar ligands.
Currently, it is challenging for FEP to transform ligands with different charges. For example, If ligand A contains a cyclohexyl substituent and it’s mutated to ligand B containing a protonated piperidine which interacts with the protein target, the change in charge leads to numerical issues preventing the calculation from giving a reliable ΔG value, so all ligands being considered are recommended to have the same charge when running a FEP experiment.
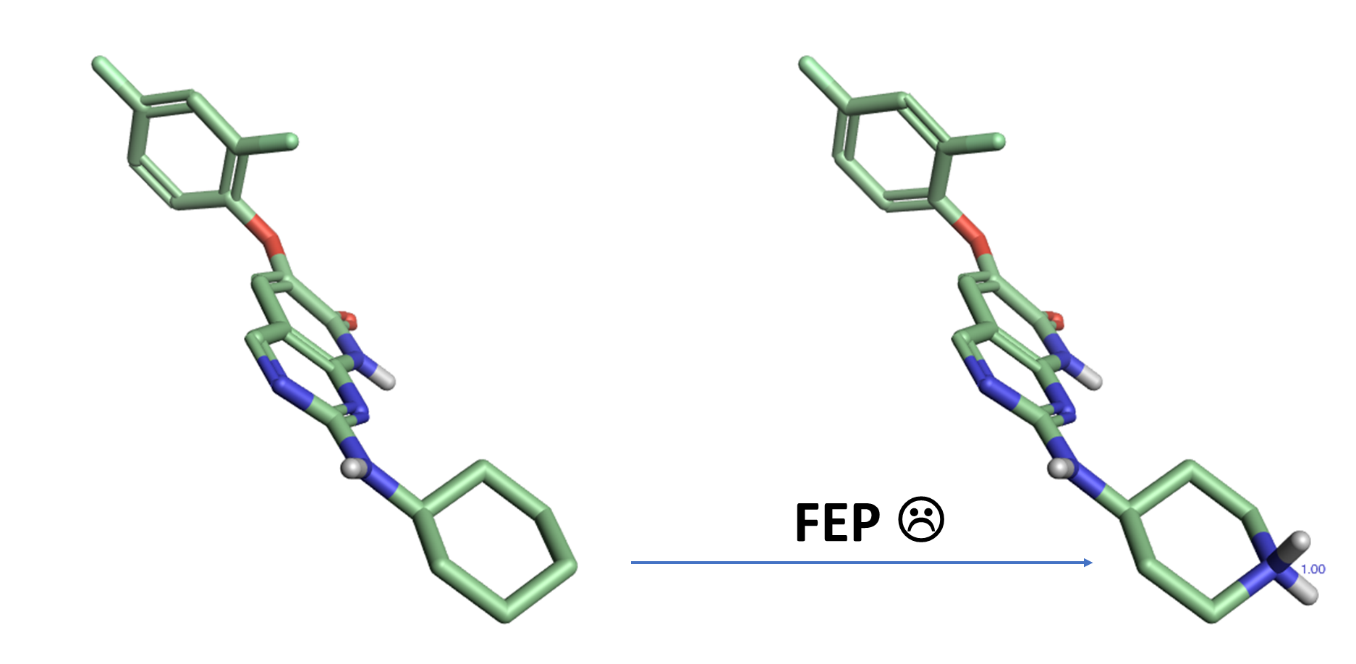
Figure 2: It is recommended that FEP calculations avoid transforming ligands with different formal charges.
Any target with a well-defined binding site could be considered for an FEP simulation. During the MD-based FEP simulation, the ligand must remain bound in order to calculate ΔG correctly. This means that binding sites which are not well defined will not give reliable results with FEP calculations. Shallow pockets as seen with many protein-protein interactions are therefore not ideally suited to FEP calculations, while small ligands such as fragments are not strong enough binders to give reliable results.
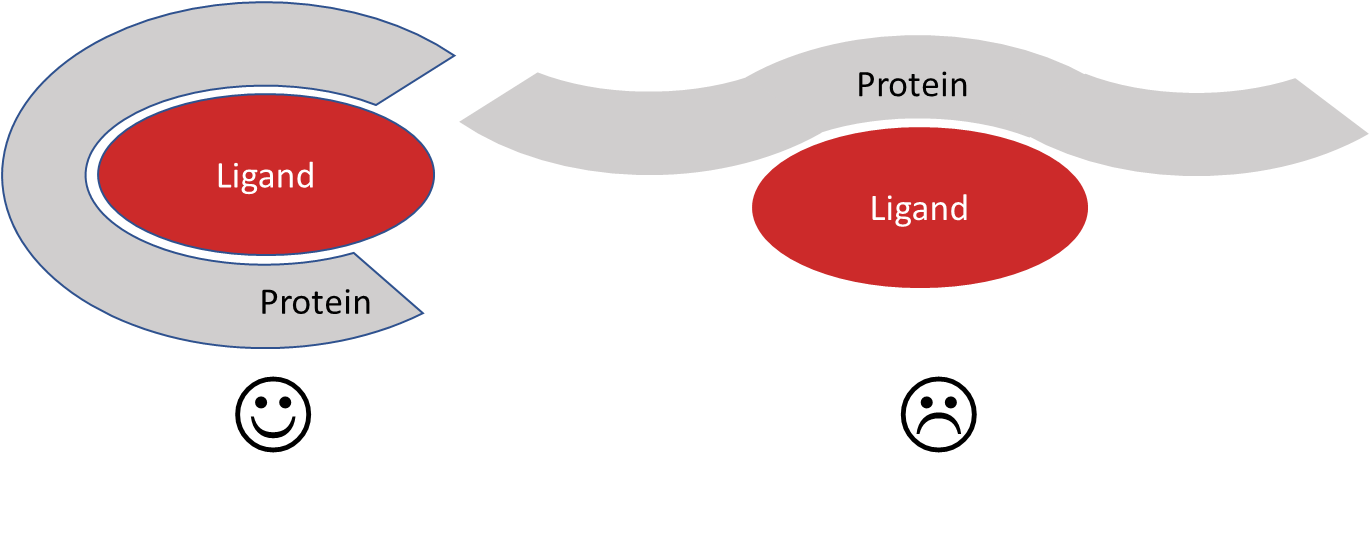
Figure 3: Binding sites amenable to FEP.
However, FEP simulations have been successfully used to help develop fragment starting points into lead-like compounds. Because it is possible to elaborate fragments using numerous different reactions and available reagents, there are many different compounds which could be potentially synthesized. Running FEP on the enumerated fragments to identify the most promising compounds will likely save time on synthesis and testing.
The sampling of system motions is also an important issue. FEP can be used to consider ligands which bind with a degree of induced fit. It has been shown with PDE2 that small side-chain movements can be adequately sampled by FEP simulations to give reliable results. In contrast, larger changes involving loop and back bone movements will not be sampled, and ligand binding requiring such large changes will not be correctly modelled using RFEB techniques. This means that ligands requiring different protein conformations are best considered in a separate experiment.
When setting up an FEP experiment, start with what is known. Firstly, prepare the protein for the simulation, making sure that it resembles the system which was used to generate the experimental results. Secondly, where possible, use a structure with a ligand bound and dock (or align) the closely related analogues into the binding site, making sure that the binding modes of the analogue scaffolds are near-identical. If docking does not give an appropriate binding mode, it may be because some induced fit is involved, and the protein structure may need to be adjusted so that the larger ligand can fit in. Prior to the FEP calculation, run an MD simulation on the bound ligand to see if it is stable – if the ligand starts to move in the MD simulations, then a correct ΔG will not be calculated in an FEP simulation, and it would be better to consider an alternative ligand as a starting point. Once the stability of the system has been checked and the ligands have been overlaid correctly, run an FEP experiment on the known ligands and ensure the correct ΔG can be calculated. When this is done, add the ligands whose binding affinity is unknown, and incorporate them into the FEP project for the ΔG calculation.
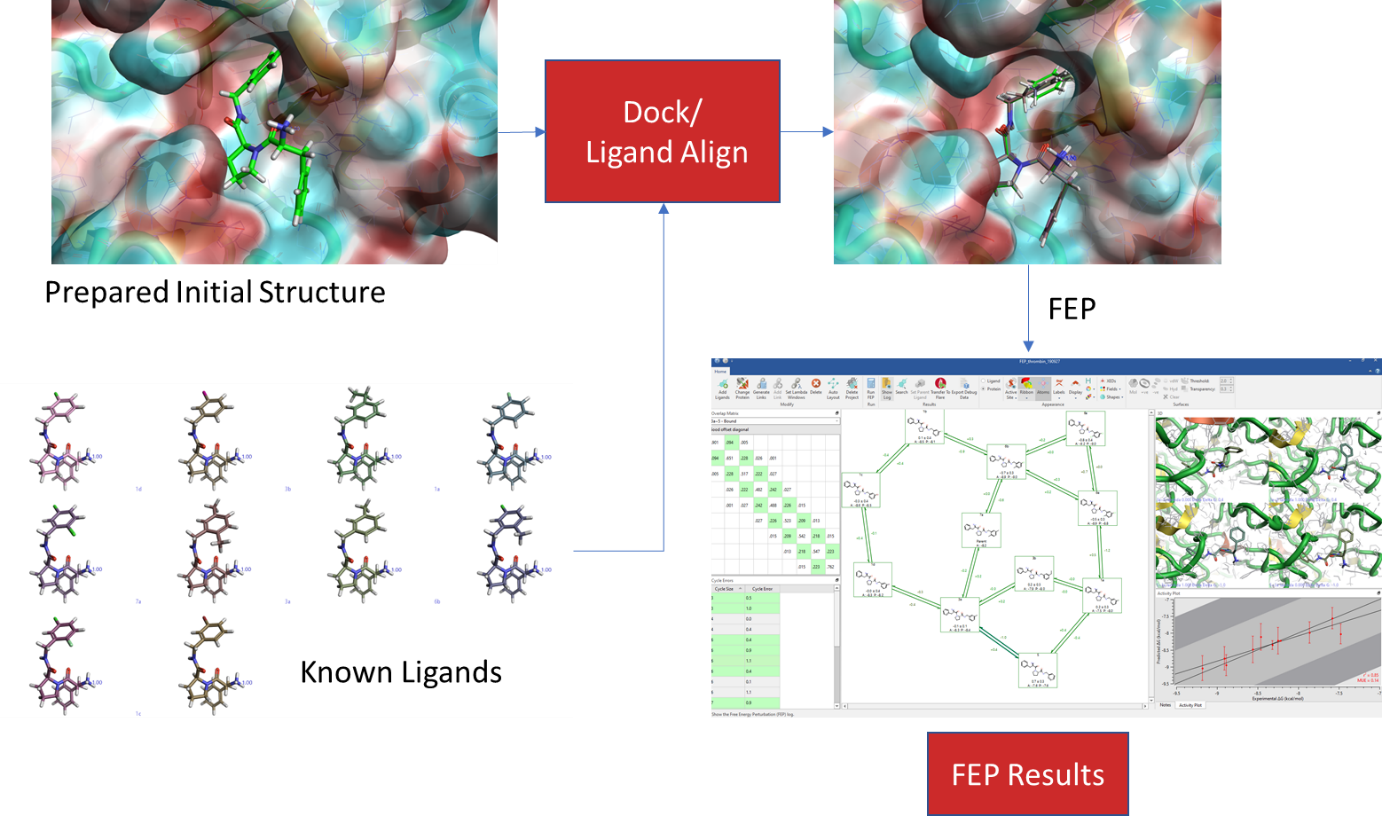
Figure 4: General Workflow for using FEP in Flare™.
One final thought. The underlying method is based on a molecular mechanics force field, and simulations are, therefore, reliant on the force field correctly describing the internal coordinates of the system as well as the non-bonded interactions between molecules. While aspects of the molecular mechanics force field describe the system well, there are some shortcomings such as inadequately describing non-classical hydrogen bonds and pi-pi interactions. It is possible to envisage a future where a QM/MM based system is used, where the ligands are described with a QM description with a surrounding polarizable MM force field environment. However, correct implementation would require some thought – especially on how the molecular orbitals between the mutating ligands would be dealt with. There are also force fields based on QM parameters, such as Parsley 3, which are being actively developed, and these newer force fields may also be useful for improving the description of the system.
Cresset released an FEP module in Flare V3 the class-leading solution for structure-based design.