Enabling Rapid Design and SAR Interpretation
The latest release of FieldAlign brings new functionality that will increase your SAR knowledge and help you design the best next synthesis. FieldAlign has always given you biologically relevant molecular comparisons that can be used to find the root causes of activity or inactivity.
FieldAlign v3.0 makes this easier by enabling sorting or filtering of your molecules using activity, physical properties or other experimental data that you have generated. Combining the filered view with FieldAlign’s excellent molecular alignments gives you a deep understanding of your compounds and the protein that they are targeting.
Using FieldAlign to design the next best synthesis target is now easier than ever. The new molecular editor enables rapid generation of iterative designs, while support for SMILES representations of molecules makes it easy to assess small virtual libraries to determine the best subset to synthesize. Improved support for copy and paste of molecules between FieldAlign, FieldStere and FieldView further simplifies the design process and makes the communication of your recommendations easy.
FieldAlign v3.0 runs on multiple CPU cores, giving you the option to use all the power of your desktop computer, or for bigger problems you can also use Cresset’s unique FieldEngines running on remote resources such as those on an in-house cluster or in the cloud to massively increase the speed of the calculations. Finally, FieldAlign v3.0 is now available as a command line binary enabling deployment in a wide variety of situations and giving ultimate flexibility in molecular alignment and scoring.
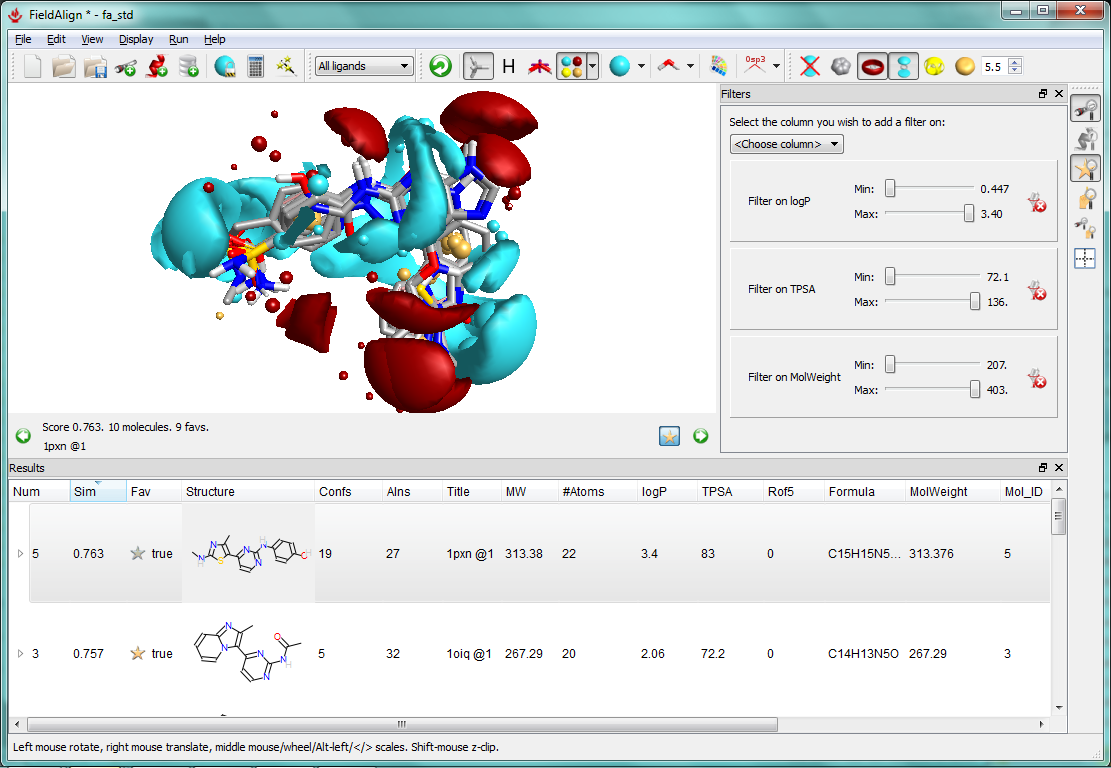
The new FieldAlign interface showing CDK2 compounds:
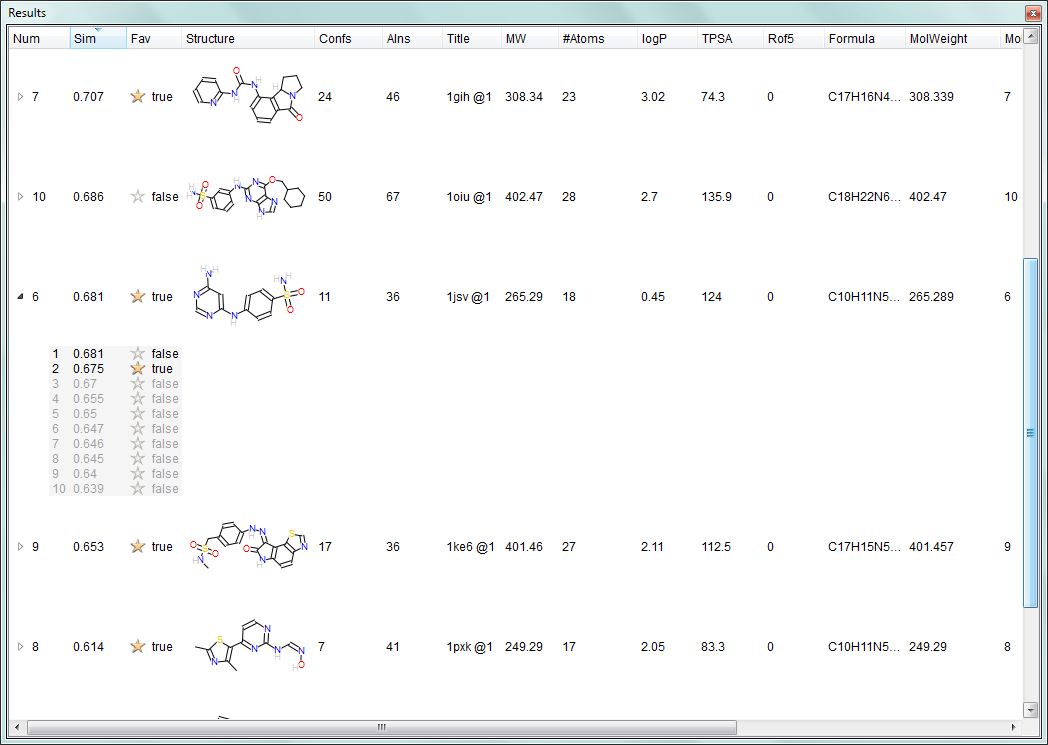
The new molecule spreadsheet in FieldAlign V3.0 showing the clustered alignment view