Improving PROTAC properties via single-point changes to linkers
We explore how computational methods can be applied to proteolysis targeting chimera (PROTAC) design, to effectively tackle some of the ...
News
As always, the UK QSAR meeting served up a smorgasbord of high-quality presentations, and lively discussion in the poster sessions given by students, academics, vendors and industrial computational chemists. There was a general focus on AI/machine learning but also a theme on free energy calculations. Below are a few of my highlights.
Jonathan Hurst gave a lighthearted and entertaining overview of machine learning, starting with the origins of machine learning referring to a paper by Hiller et al. from 1973. (DOI: 10.1016/0010-4809(73)90074-8). Reviewing progress right through to the current high impact AI publications such as Segler et al. (2018) and Ahneman et al. (2018). He asked the attendees if AI was over hyped? The majority thought it was.
Steven Oatly then attempted to challenge the audience to change their views with an excellent presentation on harnessing machine learning methods for generative chemistry design. This was coupled to an in silico scoring function (docking score) within a learning cycle, the few compounds that were selected for synthesis gave very good intrinsic activity and novel chemistry with very excited medicinal chemists. Future work would focus on how to include multiple parameters to optimize.
Richard Lonsdale presented a significant amount of work using a free energy perturbation approach for a project at GSK, with results for several hundred compounds in four distinct chemical series. Even with this volume of data there was still a need to increase the throughput, and he presented differing approaches to increase throughput from Star maps, to creating QSAR models on the predicted free energy perturbations.
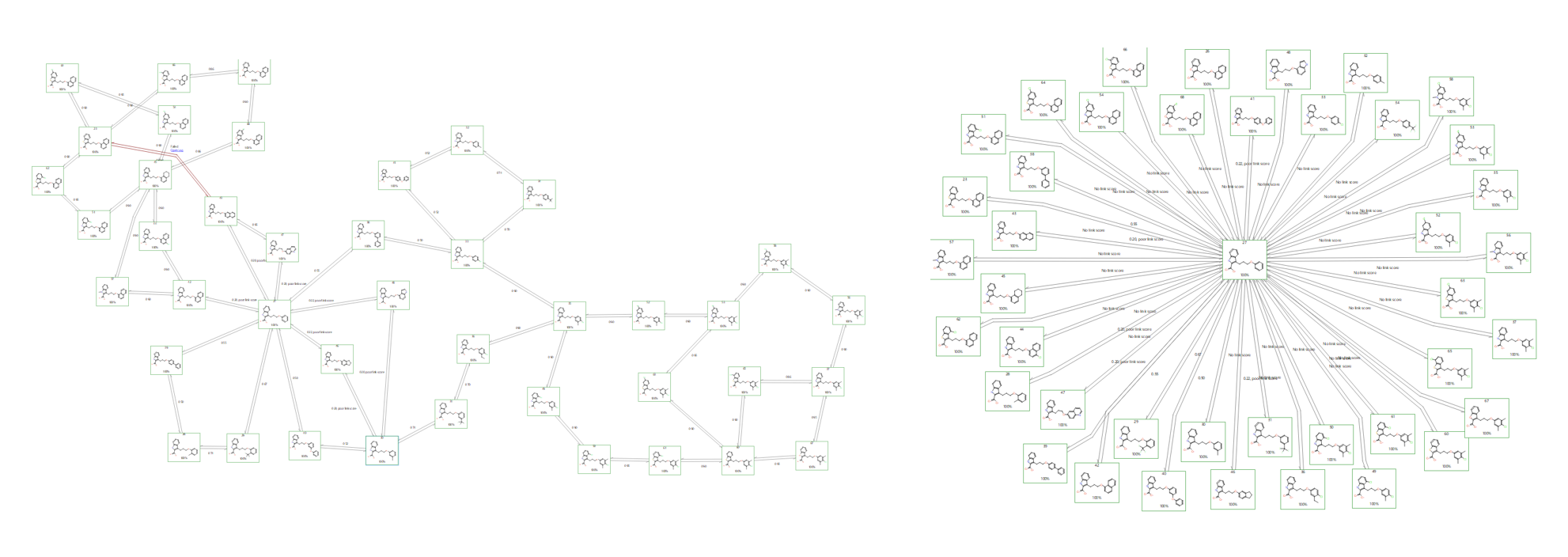 Figure 1. A traditional FEP network map (left) vs a Star map (right) created in Flare™ V3. Star maps do not contain cycles that provide validations for transitions but are generally faster to calculate.
Figure 1. A traditional FEP network map (left) vs a Star map (right) created in Flare™ V3. Star maps do not contain cycles that provide validations for transitions but are generally faster to calculate.
As an alternative to FEP, GSK’s DDU and working with Dundee University to apply Quantum Mechanics to the prediction of protein-ligand binding energies. The work is offers different advantages and disadvantages compared to MD based methods. The biggest challenge with the approach used was how to best deal with solvation, currently they use a predicted logP correction rather than an explicit solvent.
Antonella Ciancetta presented work from her time at the NIH, with a great example of the challenges of modeling proteins, which are not static structures. The target was Adenosine GPCRs and she was tasked with docking some novel chemistry selective for A3AR. Rigid protein docking failed, and so induced fit docking was required, unfortunately the results were not consistent with the project SAR. She used short MD runs to explore protein-ligand conformational space and identified that the compounds flip their pose. This new protein-ligand conformation explained the SAR and was then successfully used for further designs within the project team.
Graeme Robb presented the results of the use of molecular dynamics and monte-carlo simulations to model conformations of compounds, where a linker was designed to constrain conformational flexibility and enhance intrinsic activity. To help add in the analysis of the many conformations of each of the designs he used a variety of machine learning models, finding in his case PLS gave the most robust predictions and all methods identified additional conformational spaces that he had failed to spot manually. Ultimately whilst this work was useful for the project, Graeme felt that enthalpy was contributing to the binding energies of the compounds and the linkers would have had an impact on the electronics of the compounds too which at the time was not considered.
Figure 2. Macrocylcized ligand from PDB 5N1Z discussed in this talk.