Improving PROTAC properties via single-point changes to linkers
We explore how computational methods can be applied to proteolysis targeting chimera (PROTAC) design, to effectively tackle some of the ...
News
In a recent blog post I have shown the integration of the Jupyter QtConsole in Flare.
The Jupyter QtConsole nicely fits in with the rest of the Flare GUI and provides a comfortable Python shell environment with most of the nifty Jupyter features such as history, TAB completion, syntax highlighting, embedding of images, etc.
Since I published that post, I have started thinking that it would have been great to embed a Jupyter Notebook, as it offers an even richer interface: most importantly, it enables editing and running individual code cells, thus constituting the ideal environment for Python enthusiasts.
There were a few technical hoops that I had to jump through to get this to work, but I finally managed.
So here I am proudly presenting the Flare Python Notebook, i.e. an instance of the Jupyter Notebook embedded into the Flare GUI which has direct access to the Flare GUI objects and methods just as the Python QtConsole (Figure 1).
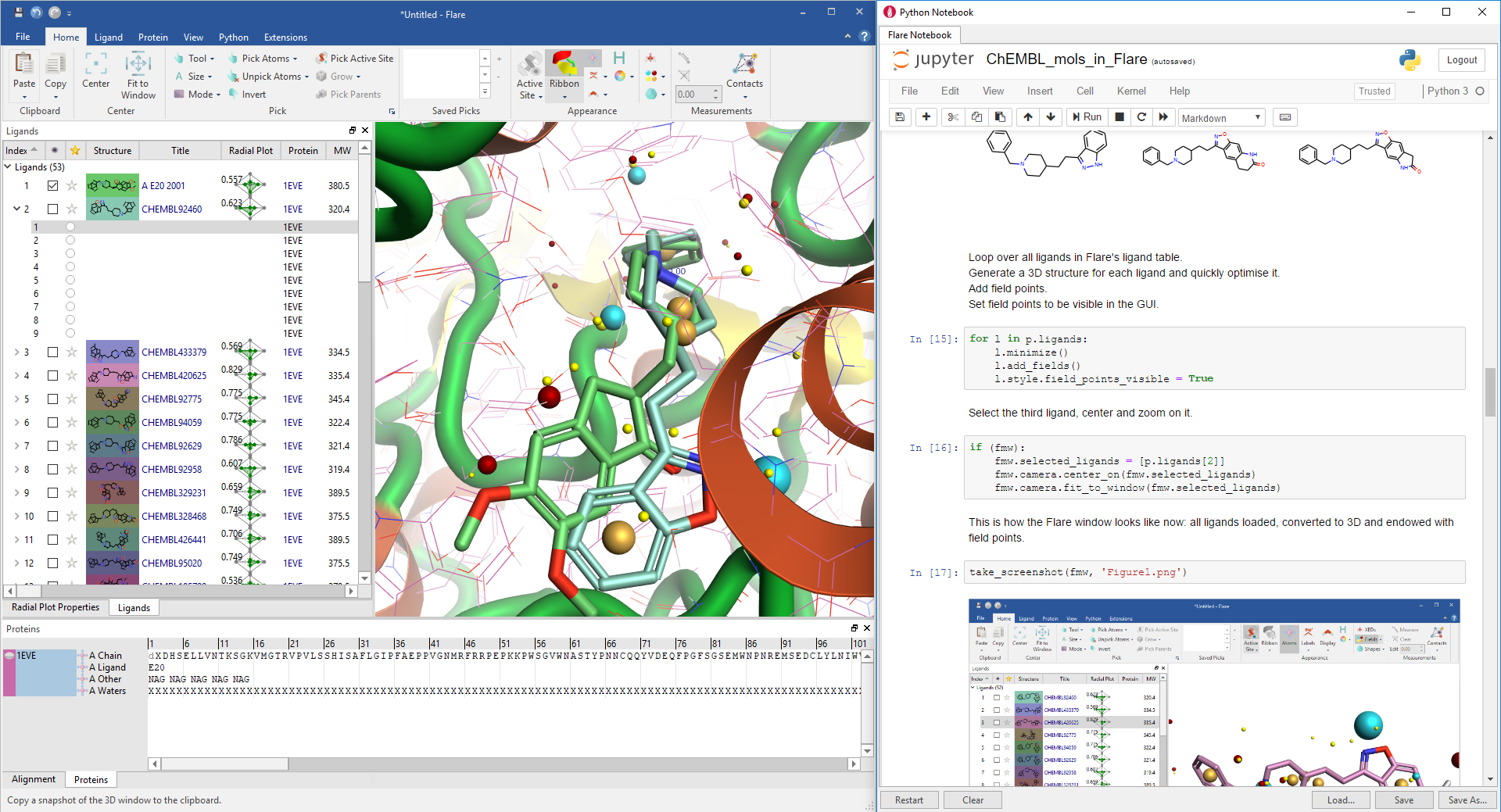
To demonstrate the new fuctionality, last week at the 7th RDKit UGM in Cambridge I gave a lightning talk showcasing a sample Python Notebook which downloads a set of AChE inhibitors from ChEMBL, loads them into Flare, generates 3D coordinates and field points, and finally docks them to a crystallographic AChE protein downloaded from the Protein Data Bank.
The RDKit is used to compute 2D similarities and maximum common substructure across ligands and to generate 2D molecule layouts. RDKit molecules are fully interoperable with Cresset molecules within Flare, so Cresset 3D technologies and RDKit methods can be synergycally combined in one Python workflow.
Matplotlib, NumPy and SciPy are used to generate a scatterplot with a regression line and compute some statistics.
The Flare Python Notebook unleashes the full potential of embedding a highly interactive Python environment within the Flare GUI.
RDKit cheminformatics methods and Cresset 3D technologies can be used side-by-side and their results visualized in real time while writing Python code, making the development cycle much more efficient and expedite.
The Jupyter Notebook has made scripting a simple everyday task for cheminformaticians, bioinformaticians and data scientists. I am confident that the Flare Python Notebook will do the same for CADD scientists and computational chemists.
And if you are not a Python guru, I can help you out; actually, I can’t wait to write my next script!
We will be releasing the Jupyter Notebook integration to our GitLab repository of Python extensions as soon as it is finalised. To find out more about Flare or to talk about how the Python integration can help you in your research or to request a Python script to achieve a particular task within Flare, then please contact us.