Improving PROTAC properties via single-point changes to linkers
We explore how computational methods can be applied to proteolysis targeting chimera (PROTAC) design, to effectively tackle some of the ...
News
Fragment-based drug discovery is an evolving area of research with unique challenges in developing the hits into lead molecules. In this case study we use the bioisostere solution, Spark™,1 to demonstrate how structure-based virtual screening effectively identifies novel InhA reductase inhibitors for tuberculosis (TB) therapies. We demonstrate how the docking score capabilities in Spark are used to grow a fragment into a new region of the active site to identify new areas of chemistry.
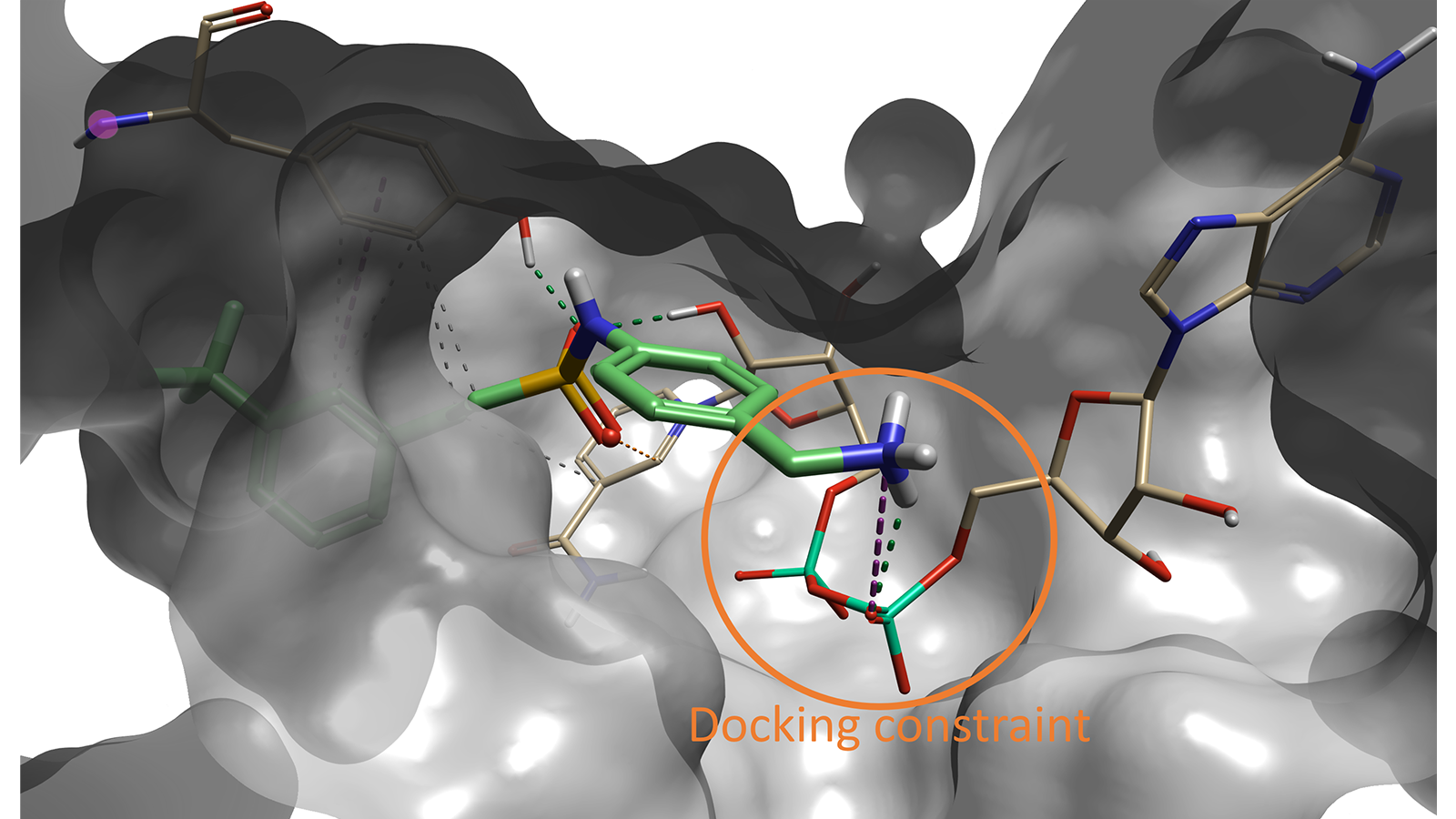
Compound #6 (out of 500) in Spark is the lead compound identified in the paper by Sabbah et al4.