Discovery
Identify and avoid problematic active sites
How druggable is your protein pocket?
Not all active sites are created equal. Even in cases where there is a lot of knowledge about a given target, companies can waste valuable development time trying to drug the undruggable. Andy Smith of Cresset Discovery shows how assessing the druggability of your active site at an early development stage can highlight any potentially costly issues of selectivity and accessibility that could slow or halt discovery progress further down the line.
Early stage project development tends to focus on whether protein ligand interactions are favourable and if so, what the opportunities are for driving activity forward. Early development is usually directed towards making compounds more active. However, a sensible approach is first to step back and examine the active site with respect to druggability.
An impartial critical analysis of the active site from the perspective of the protein, ligand, selectivity and project development can pay dividends. The important question is how to identify reasonable pockets and avoid inaccessible pockets. Can this pocket be successfully targeted by a small molecule that could progress through clinical trials and eventually become a prescription medicine? And if so, is the target selective enough compared to other members of the same protein family? Answering these questions at an early stage can avoid the cost of a project failure at the clinical stage.
Un-druggable arginase
The active site of the enzyme arginase (see Figure 1) has multiple problems which have made it notoriously undruggable. From a biological perspective, arginase is a very interesting target, yet all attempts to develop a selective drug have so far failed. Despite all the experimental and theoretical methods that have been employed by multiple companies against arginase, the results so far have only yielded a series of non-selective non-druglike compounds that contain boronic acid.
The reasons for the failure to develop arginase drugs are embedded not only in problems with the active site, but also in difficulty gaining selectivity over other members of the arginase family. The arginase active site is designed to accept only arginine and moving away from this group has proved difficult. So far, the only successful mimic for the guanidine group has been a boronic acid which binds to the metal atoms at the bottom of the pocket. Both the terminal acid and base are key for ligand binding, so generally the shape of the ligand must be very similar to arginine. There is very nearly complete homology across the arginase family, with biological in-vivo selectivity obtained by the distribution of arginase family members within the body rather than active site selectivity.
Arginase is an extreme case, but it is one that has burnt the fingers of many companies both big and small. 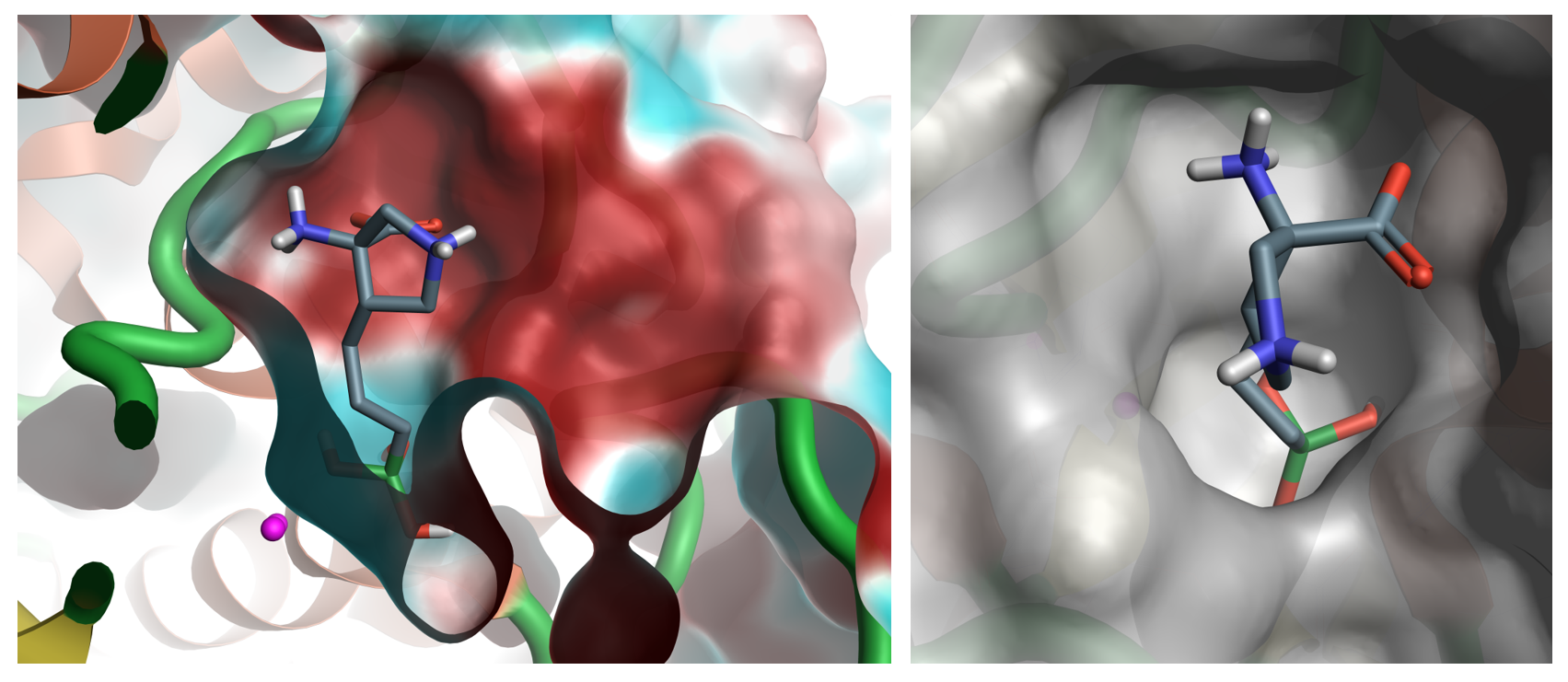
Figure 1: Arginase crystal structure 6q37 with an identified ligand in the active site. The left-hand image shows the electrostatic potential surface for the arginase active site, displaying the very polar nature of the active site. The right-hand image shows the solid surface for the arginase active site, highlighting the limited size of the active site.
How can you avoid the next arginase?
Protein ligand crystal structures are incredibly useful for projects, but they can also be blinding to the fact that some pockets are just not suitable for drug discovery.
There are multiple software packages which help to predict the druggability of protein pockets, however these tend to focus only on the character and shape of the pocket under investigation, without the wider understanding of related protein family members and problems associated with some classes of protein pockets. Instead, computational chemistry offers a different and effective way to provide a detailed assessment of druggability.
Assess selectivity across a protein family
Structural and sequence examination of protein families is a simple indication of how easily selectivity can be obtained. The structural alignment of the crystal structures for a family of protein active sites can quickly highlight those regions which can be targeted to gain selectivity and those regions which offer no opportunity for selectivity.
If crystal structures are not available for the whole family, a simple sequence alignment and point mutation analysis can give an early indication of the possibility of gaining selectivity. Full homology modeling of key off-targets may be required to fully investigate the difficulties of gaining selectivity or to gain reassurance that selectivity is possible, which may not always be apparent from a simple sequence alignment analysis process. Obviously, as the sequence, character and shape homology between active sites goes up, the difficulty of gaining selectivity usually goes up.
Assess the accessibility of a protein pocket
Examining at the size and shape of the pocket is also a useful analytical method, as small, shallow or large cavernous pockets can present insurmountable challenges for drug discovery.
Smaller compounds can be less selective as they can be weakly activity against multiple targets occupying multiple active sites. This can be a major problem as adding selectivity features can be difficult if the primary target has a small active site.
If the pocket is shallow, then gaining the necessary binding energy to make a bioactive compound can be difficult.
If the active site is very large, generating a bioactive compound can be equally hard. Designing a ligand to fully occupy the active site can lead to compounds with ADME issues. For larger sites the identification of hotspots within the pocket can help to enable the creation of small ligand by focusing the ligands to the region that are critical for protein function. Provided the hotspots are conveniently position within the active site, smaller ligand can be designed to occupy them (see Figure 2).
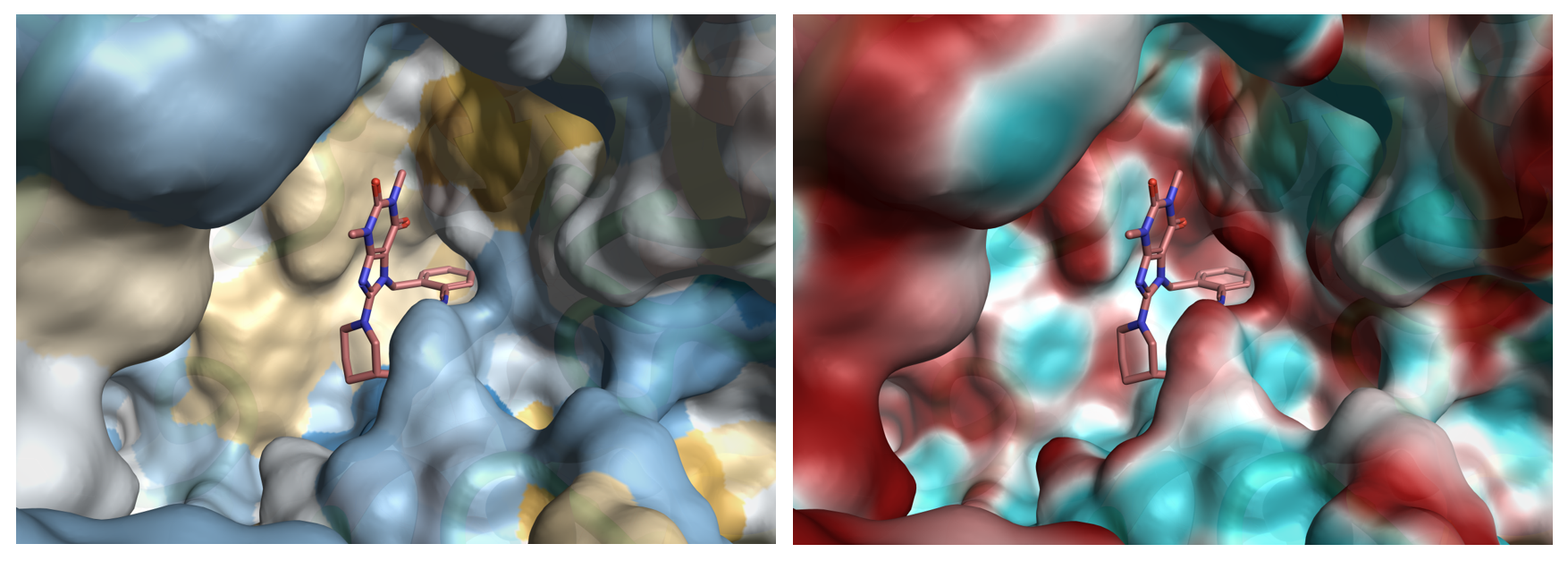
Figure 2: Hydrophobic and electrostatic potential energy surfaces for the DPP IV crystal structure 3g0d. Although the active site is fairly large, key hotspots have been identified. The two surfaces show the mixed nature of the active site which helps to design balanced ligands with improved physiochemical and ADMET properties.
Examine the natural substrate
The natural substrate for an active site can also be an indication of druggability as this is the ligand we are going to be competing against. Highly charged ligands like those containing phosphate groups present a difficult target as the designed ligand may need to compete with a phosphate group. A highly charged ligand tends to be required in order to compete with phosphate, but these types of systems tend to have ADME issues. On the other hand, systems with lower charge have a better ADME profile but cannot compete with phosphate binding.
There are examples of bioisoteres which have successfully worked in a phosphate binding region, but these types of active sites tend to be troublesome. Active sites which bind large, floppy, lipophilic groups are also problematic as they tend to have multiple diffuse lipophilic interactions and designing a competitive molecule tend to move the molecule into very non-druglike property space.
Identify and avoid problematic active sites
By combining analyses from the perspective of protein, ligands and general medicinal chemical knowledge it is possible to differentiate between pockets which are going to be problematic and those which are going to be less troublesome to develop.
Even if your protein has a difficult active site, it may still be possible to target it or even approach the project from a different perspective and identify an allosteric binding site which avoids the problems of the primary active site.
If you are concerned that you are working with a troublesome active site, contact us for a free confidential discussion to see how we can help progress your project.
Discovery
Contact us for a free confidential discussion
We help you reach your next milestone faster and more cost effectively
Contact us for a free confidential discussion