Discovery
You have a crystal structure, but what do you do with it and how do you use it?
The increasing availability of protein crystal structures continues to have a huge effect on drug discovery, improving the ability to recognize key protein ligand interactions. The ability to generate bespoke crystallographic data on a project can highlight additional chemistry opportunities that can propel a project forward in new directions. However, like all data, the information must be processed and analyzed to ensure the reliability of the structure and that the maximum information is obtained. This process is covered by the general method of protein preparation.
The importance of protein preparation
The goal of protein preparation is to generate one or more protein models that represent the bioactive conformation(s) of the protein when ligands are bound. This can be a single representation, but it is more likely to be an ensemble of closely related structures showing the concerted movements of protein and ligand upon binding.
To release the full potential of crystallography, careful structural preparation is vital. A badly prepared structure can lead to an unrealistic structure, misinterpretation of results and can retard project development. Badly prepared structures can lead to confusing interaction analysis, incorrect assignment of binding mode and in general can greatly reduce the value of crystallographic input into a project.
There are multiple tools which automate the process of protein preparation, such as Flare™, but these cannot be used successfully without critical evaluation of the results. Blindly relying on automated tools without experienced evaluation is very likely to produce less than optimum outcome for project development and degrade the information that can be extracted from crystal structures.
Assess your starting point
The first stage of protein preparation is to look at the reported data to get an estimation of the quality of the data and how much trust can be placed in the crystal structure. When available, electron density maps provide greater understanding of the crystallographic quality and a better understanding of the unresolved regions in the crystal structures. Whether water molecules are strongly or weakly resolved can be especially important in older crystal structures when water was sometimes used to in improve the reported resolution without justification from the electron density.
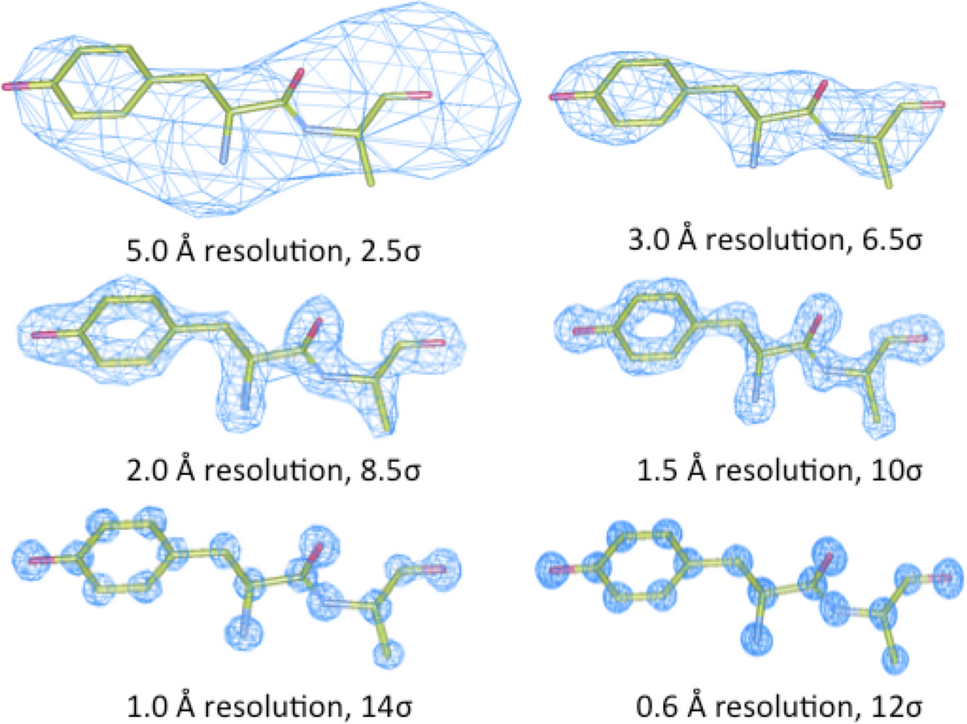
Figure 1: Different electron density maps at differing resolutions1.
Electron density analysis ranges from quickly examining the electron density map to assess whether it’s well defined or merely a general smear of electron density, right the way through to potentially rebuilding part of or even the whole structure in particularly bad cases. Relating the structure to the electron density is both informative and essential for a full understanding of the quality and key features of a crystal structure. The program WinCoot2 which is freely available can be used by both experienced users and those new to the electron density analysis to glean additional information.
Investigation of the unit cell is also enlightening as it can highlight regions of the protein which are potentially deformed due to crystal packing forces. If additional crystal structures are available these should also be compared as different crystal structures can have different unit cell and so different crystal packing.
Decide what to do with unresolved regions
Once familiarity with the structure and the electron density map has been obtained, this can be used to make informed decisions around regions of the protein that are fully or partially unresolved, including alternate side chain conformations and water molecule positions. When a residue side chain exhibits multiple conformations close to the ligand binding site, this leads to the situation where multiple models are required to describe the protein and ligand binding modes. Only by understanding the underlying electron density maps is it possible to be fully confident in describing these alternative forms. The use of Ramachandran plots allows a very quick identification of protein regions which are atypical and may require investigation and correction.
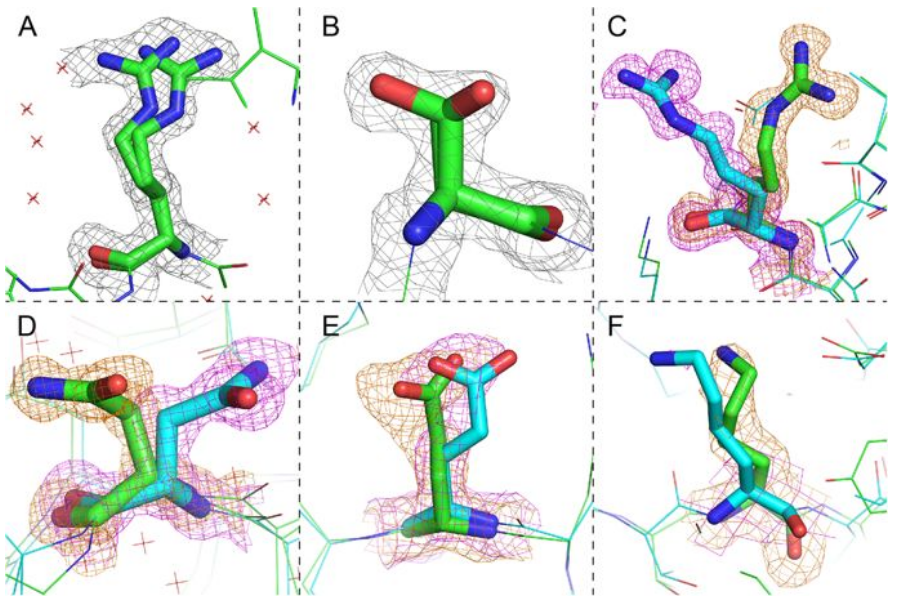
Figure 2: Electron density maps for residues with alternate sidechain conformations3.
Identify key structural features
Examination of the structure for potential disulphide bonds, salt bridges and metal binding sites should be carried out once there is confidence in the rough positions of the residue side chains. These interactions can be key structural features stabilising and defining the protein tertiary structure. Very careful investigation and assignment of these features is required as an incorrect assignment can have a negative effect on the quality of the models, resulting in an unrealistic protein structure.
Disulphide bonds are reasonably simple to identify due to the proximity of the cysteine residues, but care must still be taken to ensure that the disulphide bond is real. Salt bridges are also reasonably easy to spot due to the proximity of the charged residue side chains and their geometry, difficulties can occur when the resolution is low and side chains are not fully resolved, but a strong salt bridge should be more resolved than generally disordered side chains. Metal binding sites pose a problem as we need to know the oxidation state of the metal to assign the correct coordination geometry. As metals in proteins tend to exhibit multiple oxidation states, with coordination sites possibly occupied by water molecules, very careful analysis is required - on a case by case basis.
If the crystal structure has a ligand bound at a minimum, any residue within six angstroms should be manually inspected to investigate ligand interactions and structural deformations resulting from ligand binding.
Assign hydrogen atoms and protonation states
Once you are confident with the overall protein structure, hydrogen atoms and protonation states can be assigned. Usually this is automated within a software package but, as with all automated processes, this must be checked. This is especially true around any ligands. Ligands can change the typical nature of protein amino acids and ligands tend not always to be correctly protonated, so careful analysis and critical assessment is vital.
Protonation software frequently provides incorrect assignment for the following problem residues: cysteine, histidine, asparagine and glutamine, so these residues require special attention. Cysteine residues should be checked for any missed disulphide bonds. A cluster of cysteine residues can be indicative of a metal binding site and, depending on the environment, a cysteine residue can sometimes be de-protonated, especially when it is in an active site and part of a catalytic cycle. Histidine has two tautomeric forms which may have different preferred rotational orientations, and it may be protonated depending on its local environment. This means that there are potentially six conformations that a histidine can adopt for a single resolved side chain position. Finally, the side chain oxygen and nitrogen atoms of asparagine and glutamine are difficult to resolve crystallographically, therefore examination of the environments around these residues must be carried out to correctly assign their orientation. This can be especially difficult or impossible if there are water mediated hydrogen bonds.
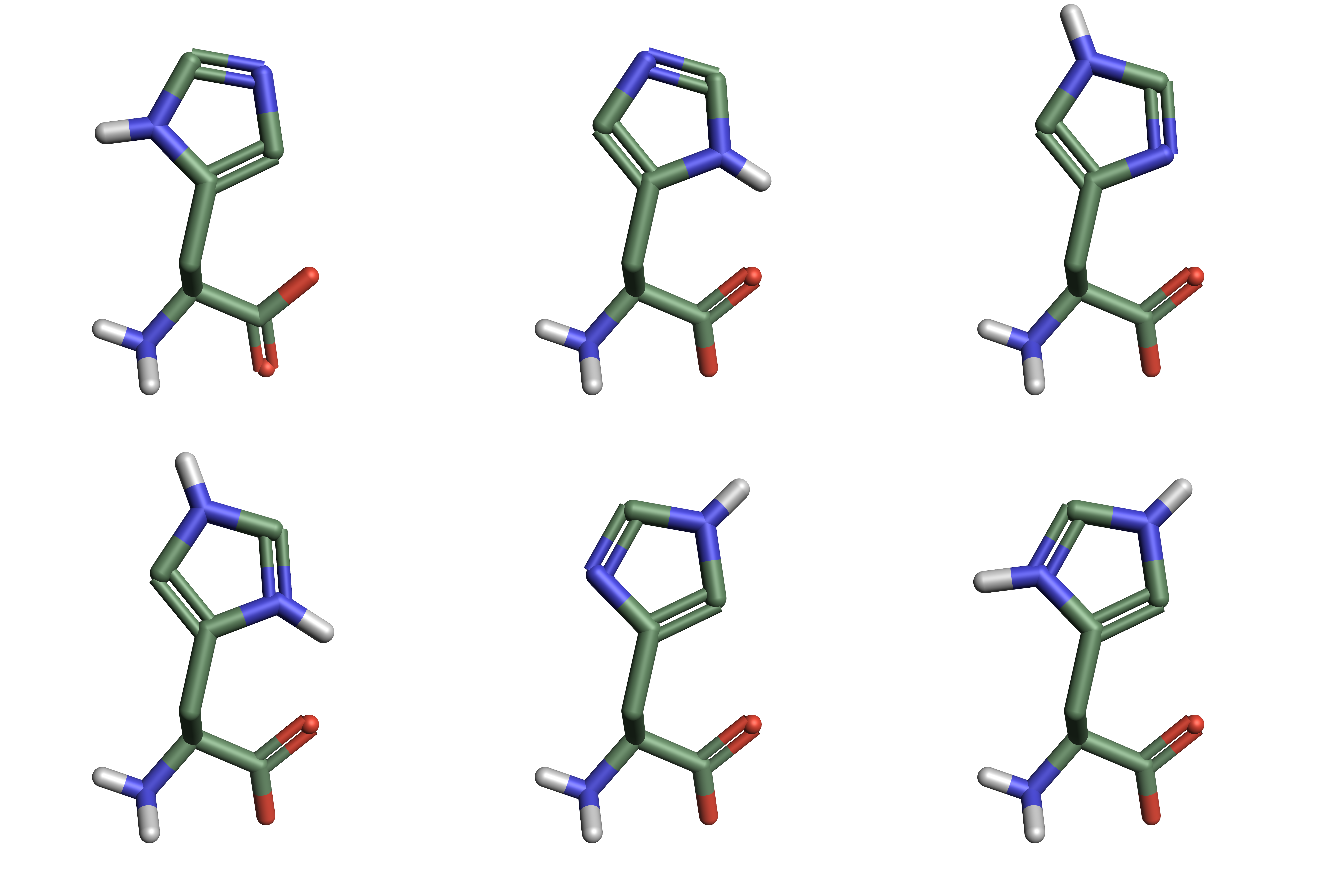
Figure 3: Tautomeric and Rotational forms of histidine.
Geometry optimization
When confident that residues are correctly assigned, and the protonation states are correct, the structure has been thoroughly checked and the Ramachandran plot is acceptable then the structure is ready for geometry optimization. A series of energy minimisation should be undertaken to test the overall stability of the protein. Typically, this is a multi-step approach gradually releasing more and more of the protein as described in the workflow below:
- Free up the hydrogen atoms to allow for the relaxation of hydrogen bonds
- Carry out side chain relaxation to allow side chain clashes to reduce while the backbone is held rigid preserving the tertiary and secondary structures
- Carry out full protein minimization to allow for the general relaxation of the whole protein
Regions which display a large amount of deformation should be revaluated and the reason for the structural change established and rectified if required. The energy-minimized structure should be related back to the electron density map to confirm that the structure is still representative of the observed electron density and conforms to the acceptable regions within the Ramachandran plot.
Once a stable structure is obtained from energy minimization, this system may be investigated using molecular dynamics to assess its stability over time. The coming release of Flare this functionality has been added to enhance the analysis of protein structures Once again the structures observed in the dynamics should be related back to the electron density map and any region that shows a large degree of deformation should be investigated to establish the cause and rectified if required.
The result of this process should be a protein crystal structure in a low energy conformation that is consistent with the observed electron density map and that is suitable for structure-based drug design. The protonation states of any ligand and residues in the active site should be understood and potential transformation in and around the active site should also be identified. Comparisons of the residue RMSD during the molecular dynamics to the observed electron density maps to confirm they are consistent. The molecular dynamics can also help to identify those water molecules which are truly structural from those which are transient as understanding the position of water molecules in the protein can often help us to understand the protein. Using the 3D-RISM protocol within Flare can help to confirm the stability of those water which appear positionally conserved during the molecular dynamics calculation4.
Free confidential discussion
Contact Cresset Discovery for a free confidential discussion to see how we can help you enhance your protein preparation process.
References
- High-Resolution Crystal Structures of Protein Helices Reconciled with Three-Centered Hydrogen Bonds and Multipole Electrostatics, Daniel J. Kuster, Chengyu Liu, Zheng Fang, Jay W. Ponder, Garland R. Marshall 2015 PLOS ONE https://doi.org/10.1371/journal.pone.0123146
- WinCoot http://bernhardcl.github.io/coot/wincoot-download.html
- Quantifying side-chain conformational variations in protein structure, Zhichao Miao, Yang Cao, 2016 Vol 6 Page 37024 https://doi.org/10.1038/srep37024
- Putting-electrostatics-and-water-at-the-center-of-structure-based-drug-design https://www.cresset-group.com/media/uploads/files/
Discovery
Contact us for a free confidential discussion
We help you reach your next milestone faster and more cost effectively
Contact us for a free confidential discussion