Discovery CRO
Virtual screening
Dramatically increase your in vitro screening hit rate, at a fraction of the cost, with diverse new structures utilizing 3D electrostatic and shape based screening
In silico screening is a cost-effective approach to complement and increase efficiency of your in vitro screening, allowing exploration of larger chemical space, discovering diverse chemotypes and generating new IP in the search for novel hits. Relying on the simple premise that a bioactive conformation encodes, in its electrostatic field and shape, recognition features optimized for interaction with its protein target; compounds with higher similarity to the bioactive conformation, having a higher probability of being active, are identified.
Cresset Discovery CRO's computational scientists collaborate with your team to apply solutions powered by Cresset field technology to select new chemical matter, ensuring the best compounds are submitted for screening.
Enumerated libraries
3D screening of commercially available,
off the shelf, compounds
Rapid exploration of millions of compounds
Combinatorial spaces
3D screening of ultra-large chemical spaces
Fast screening of billions of compounds
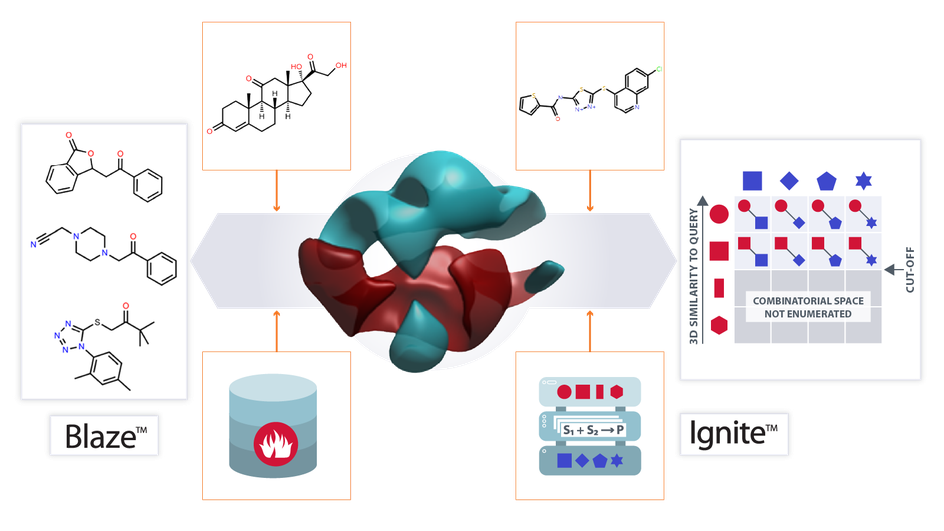
Access comprehensive sampling of chemical space and receive a rank ordering of compounds that can be used as a means of selection and prioritization
Enumerated libraries
Through the skilled application of Cresset’s proprietary virtual screening software platform, Blaze™ our Discovery CRO team uses the 3D electrostatics and shape of a defined ligand to rapidly search large chemical collections for molecules with similar properties. By adjusting the level of influence played by each of these factors, our team can achieve a high diversity of hits, with hit rates as high as 30%.
The screening databases available through the Cresset Blaze server contain more than 30 million compounds. These are molecules from easily accessible vendor databases.
Blaze performs a comprehensive sampling of chemical space, providing novel lead-like structures attributed to Cresset’s field technology, without discriminating based on substructure and chemotype similarity.
Our scientists use their expert insight and experience to select the optimal criteria for the screen and to interpret and elucidate the results, proposing a final selection of hits to progress.
Learn more about Blaze™ platform.
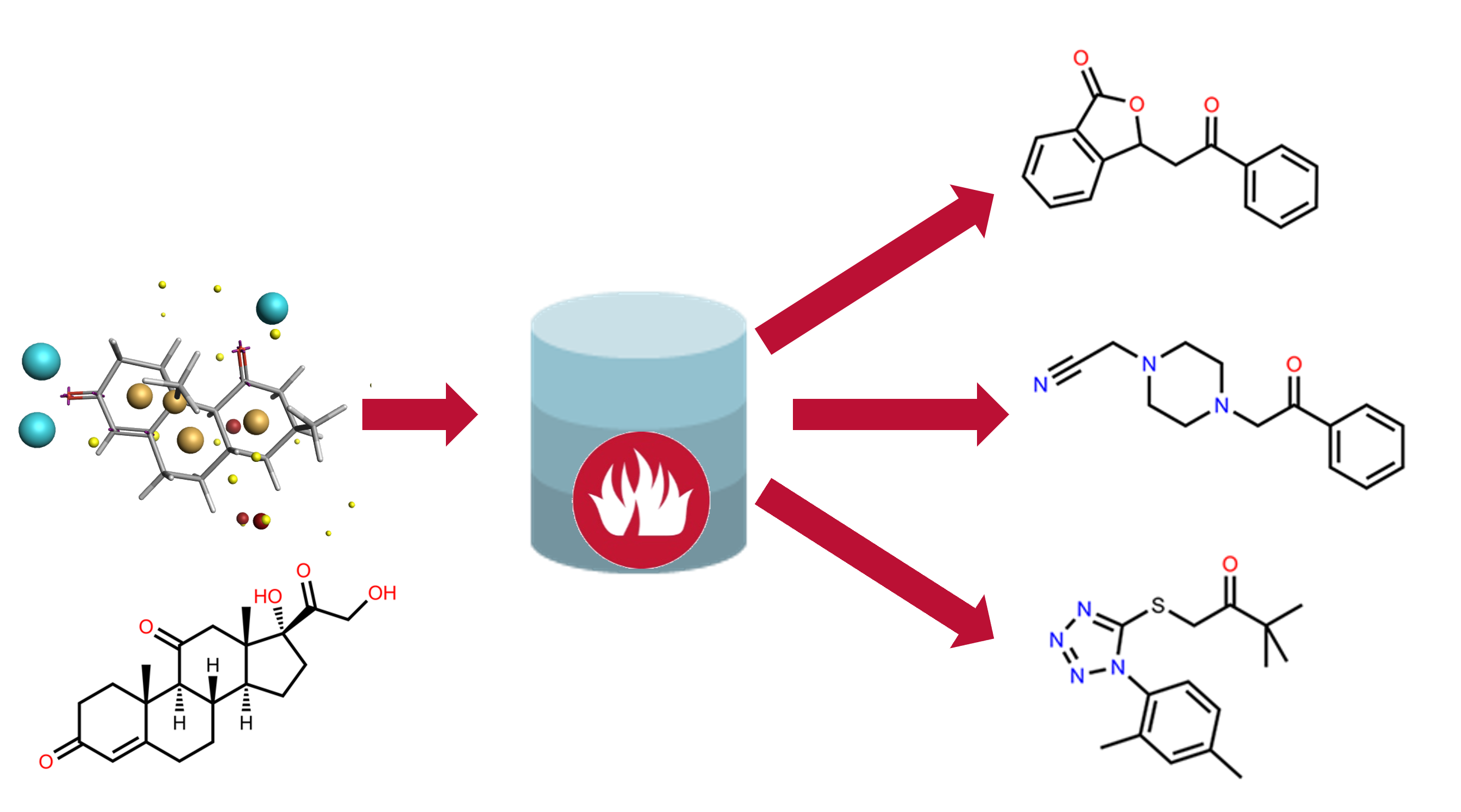
Field point representation as the 'template' for Blaze virtual screening
Combinatorial spaces
Ignite™ is a new technology to enable rapid 3D virtual screening in combinatorial chemical spaces. Conventional virtual screening on large data sets is typically carried out by comparing 2D molecular fingerprints. This makes screening relatively fast but has the drawback that top-scoring results are less likely to be actual hits.
Synthesizable chemical spaces are vast and routinely contain billion of molecules resulting in the fully enumerated libraries practically impossible to search. Ignite utilises the combinatorial nature of most chemical spaces i.e., a relatively small number of building blocks/fragments are combined to create a much larger number of enumerated compounds.
The Ignite algorithm performs a fast initial screen in reagent space. Reagents for each virtual library are converted into ‘product-like’ fragments, converting their reacting functional groups into minimal representations of the library products. These product-like fragments are then screened against the query molecule to determine which ones are likely to provide significant shape and electrostatic similarity.
Watch our webinar demonstrating the application of Ignite™ technology to Enamine's REAL Space.
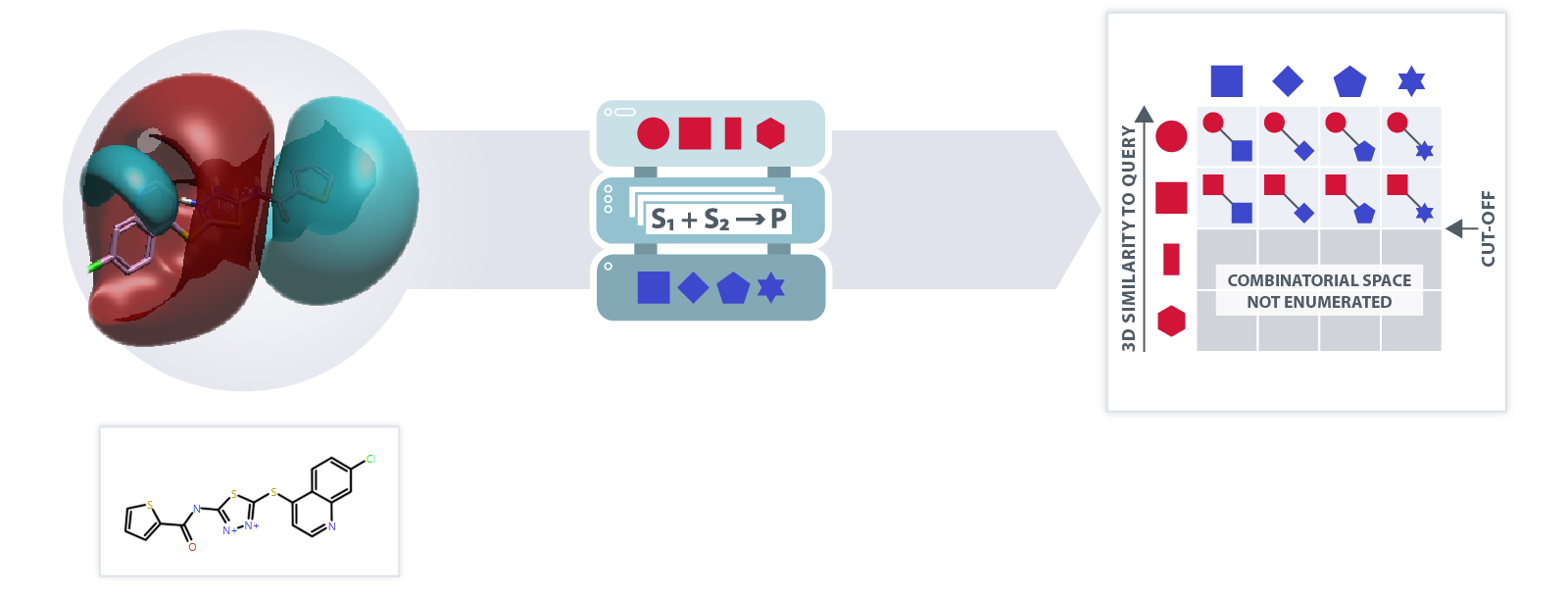
Fast 3D virtual screening of billions of compounds with relatively modest CPU resources. Exploration of ultra-large chemical spaces in terms of synthons and reactions with rapid screening of large parts of the space.
Cresset Discovery CRO has delivered hundreds of successful virtual screening projects. We work with you to devise the best approach for your project through:
- The scientific knowledge, experience and sophisticated methods required to set up and run experiments, and to optimize and interpret the results to get maximum output
- The computing infrastructure needed to run and manage a high volume of experiments in parallel, allowing you to easily scale up operations and advance your project
Read our case studies.
Contact us for a confidential discussion.
Contact us for a free confidential discussion
We help you reach your next milestone faster and more cost effectively
Contact us for a free confidential discussion