flare™
QSAR models
Decipher complex SAR and choose the best molecules to make
Decipher Structure-Activity Relationships
The Activity Atlas™ and Activity Miner™ components of Flare enable you to understand the electrostatics and shape features which rule the SAR of your ligand series.
Summarize SAR
Activity Atlas visually summarizes the SAR for your ligand series generating informative 3D maps based on a Bayesian analysis of the:
- Average electrostatic and shape character of active molecules
- Activity cliffs information from the Activity Miner disparity matrix
- Electrostatic and shape regions that have been explored so far
Activity Atlas is particularly useful for those project teams where there is not enough SAR for building a quantitative SAR model, and to aid the interpretation of quantitative Machine Learning models.
When viewed in combination with 3D maps of protein electrostatics, Activity Atlas maps are a powerful way to get an in depth understanding of the SAR for your ligand series.
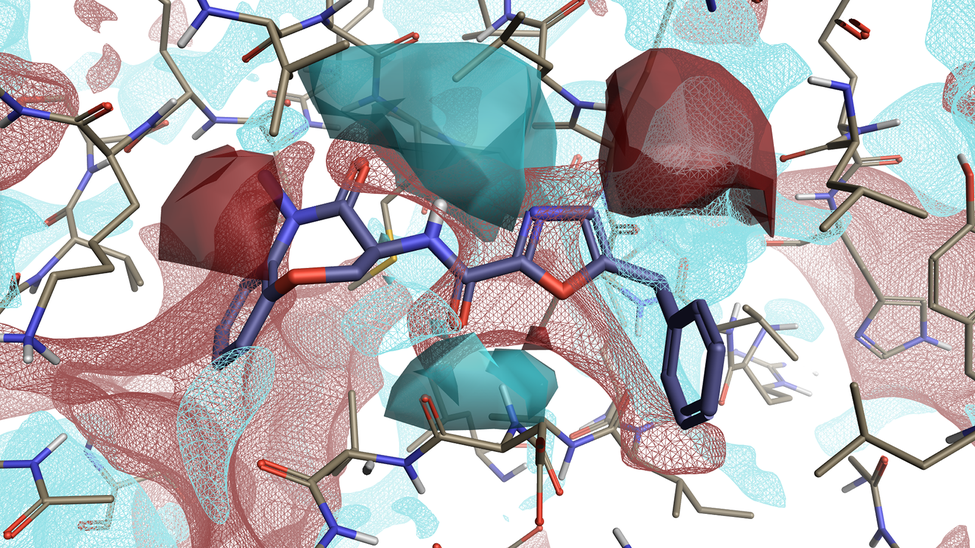
Activity Atlas visually summarizes the SAR for your ligand series into informative 3D maps which viewed in combination with 3D maps of protein electrostatics are a powerful way to get an in depth understanding of the SAR for your ligand series.
Rapidly navigate complex SAR
Activity Miner enables the rapid navigation of complex SAR by analyzing activity and selectivity cliffs to highlight critical regions in the SAR landscape where major changes take place.
Multiple views of your data help you find key molecule pairs in your SAR where an activity cliff is observed. For each pair, Activity Miner shows you how the electrostatic and shape properties differ, building an understanding of how to design better compounds with better properties.
The different views enable you to focus on different aspects of your SAR.
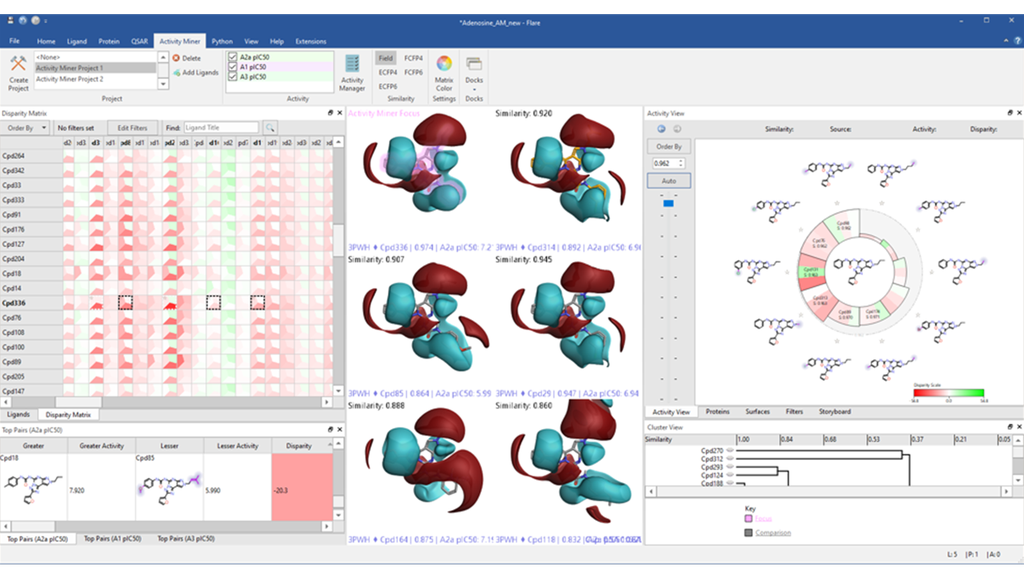
Find and display activity cliffs using multiple activities across all compounds or focus on only those present for a single compound pair.
Robust QSAR models for activity prediction
Build a wide range of quantitative models to predict the activity and ADMET properties of new compounds and prioritize the best molecules to make.
Field QSAR: Prediction and interpretation
Field QSAR models provide a global view of your SAR data. They work well where the SAR landscape is smooth – small changes in a molecule lead to small changes in activity. Where a robust model is obtained it can be usefully employed in the prediction of activity for new molecule designs, aiding the prioritization for synthesis. Alongside each prediction, the fit of a particular compound to the model can be studied to understand what is favorable or unfavorable about the design. This gives feedback to improve the design but also aids in deciphering the model and the reason a particular compound is highly active or inactive.
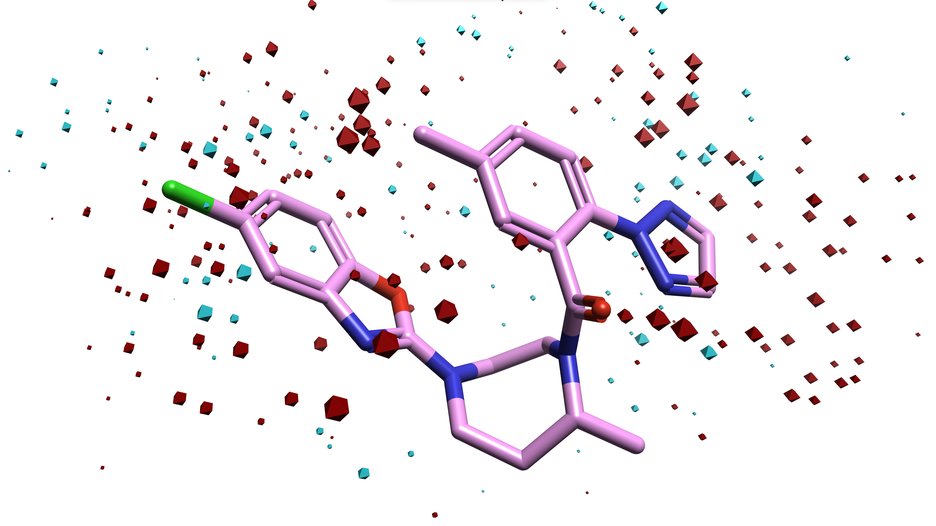
Field QSAR models provide a global view of your SAR data, to give you both prediction and interpretation.
Robust Machine Learning models for predicting activity and ADMET properties
Build predictive Quantitative SAR models by choosing among several robust and well validated machine learning methods – or by running them all and letting Flare pick the best.
Calculate quantitative models of regression – suitable when the biological activity data are real values such as pKi or pIC50 – or classification, to model qualitative biological data or data expressed as activity ranges (e.g., % inhibition).
Predict the activity or category for new compounds using consensus QSAR, to prioritize the molecules all models agree would be a good idea to make.
In Flare, predictive QSAR models for activity and relevant ADMET properties can be calculated using a variety of descriptors:
- Flare’s 3D descriptors, modeling the shape and electrostatic character of aligned molecules
- Fingerprints and descriptors from the RDKit
- Custom 3D/2D descriptors.
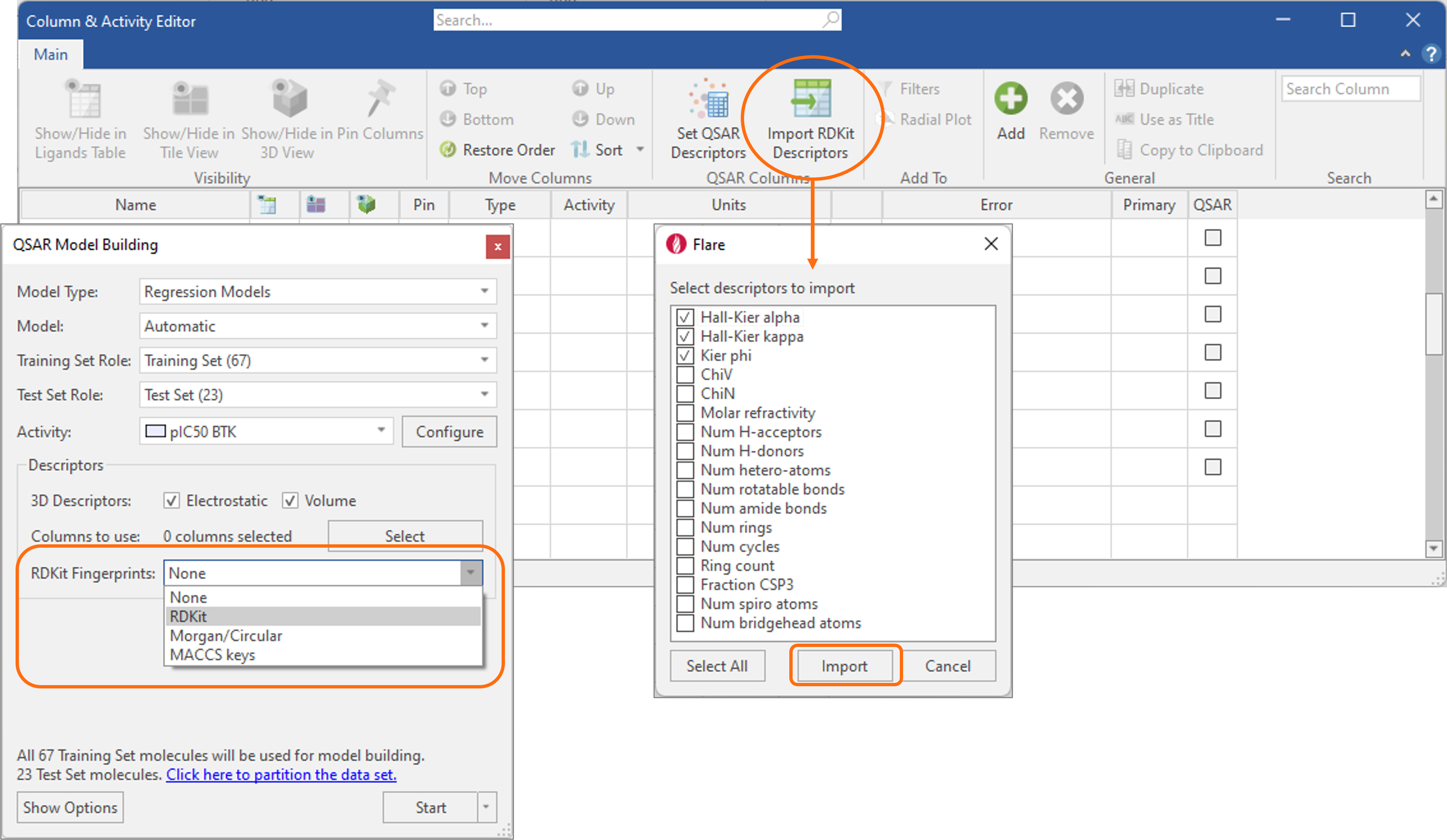
Build predictive QSAR models of activity and ADMET properties using a variety of descriptors.