Anand Balupuri†, Giovanna Tedesco†
†Cresset, New Cambridge House, Bassingbourn Road, Litlington, Cambridgeshire, SG8 0SS, UK
Abstract
Metabolic stability of a compound is a key factor in drug discovery and development. Several compounds with promising pharmacological properties fail to become drugs due to poor metabolic stability profiles. Bioisosteric replacement is a powerful tool for modulating properties of the compounds. In this case study, Spark1, Cresset’s bioisostere replacement and scaffold hopping software is used to demonstrate how bioisosteric replacements can be useful in addressing metabolic liabilities.
Introduction
The mitogen-activated protein kinase (MAPK) pathway is a major cell-signaling pathway and aberrant activation of the pathway frequently drives tumor growth.2 The extracellular signal-regulated kinase 1 and 2 (ERK1/2) play a central role in the MAPK pathway3 and are promising target for the treatment of tumours.
Various ERK1/2 inhibitors with promising preclinical efficacy have progressed into clinical trials in the last few years.4 The isoindolinones are a novel class of ERK1/2 inhibitors recently reported by Astex Pharmaceuticals.5 Compound 7 (Figure 1 – left and Table 1) is a highly potent and selective ERK1/2 inhibitor, shows promising pharmacokinetics profile in mice, however, has a short in vitro half-life (5.1 min) due to oxidative metabolism by Cytochrome P450 3A4 (CYP3A4).6 Identification of metabolites revealed that both ends of the compound 7 (the tolyl methyl group and the oxan ring) undergo significant oxidative metabolism.
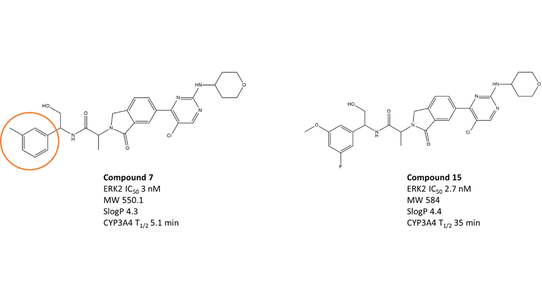
Figure 1. Left: Compound 7 is a highly potent and selective ERK1/2 inhibitor with a short half-life. Right: The optimized Compound 15 shows enhanced potency and a longer half-life.
Lead optimization strategy to enhance the metabolic stability of Compound 7 included direct blocking of the vulnerable sites by replacement with more stable isosteres (SAR data for analogues is shown in Table 1), conformational rigidification, and modulation of the overall physicochemical properties of the compounds. These efforts led to the identification of compound 15 (Figure 1 – right and Table 1), showing slightly higher potency and significantly improved metabolic stability against CYP3A4, with in vitro half-life of 35 min. Compound 15 (ASTX029) is currently in a phase I−II clinical trial in patients with advanced solid malignancies.6
Table 1. SAR exploration for the tolyl moiety replacements.6
| Compound | R | ERK2 IC50 (nM) | CYP3A4 T1/2 (min) |
| 7 | 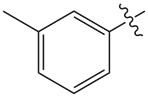 |
3.0 | 5.1 |
| 14 | 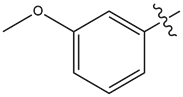 |
1.3 | 5.5 |
| 15 | 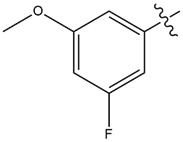 |
2.7 | 35 |
| 16 | 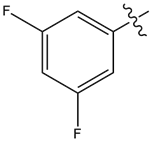 |
43%@1 | <5 |
| 17 | 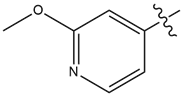 |
1.8 | 15 |
The published X-ray crystal structure of lead compound 7 in complex with ERK2 (PDB code: 6G9N) shows that this molecule occupies both the hinge region and the secondary pocket (Figure 2), which is considered to be required to elicit the desired dual pharmacological profile.6 The tolyl moiety binds to the secondary pocket and exhibits an aromatic-aromatic interaction with TYR 64. The oxan ring binds the nearby hinge region and does not display any specific interaction with the active site of ERK2.
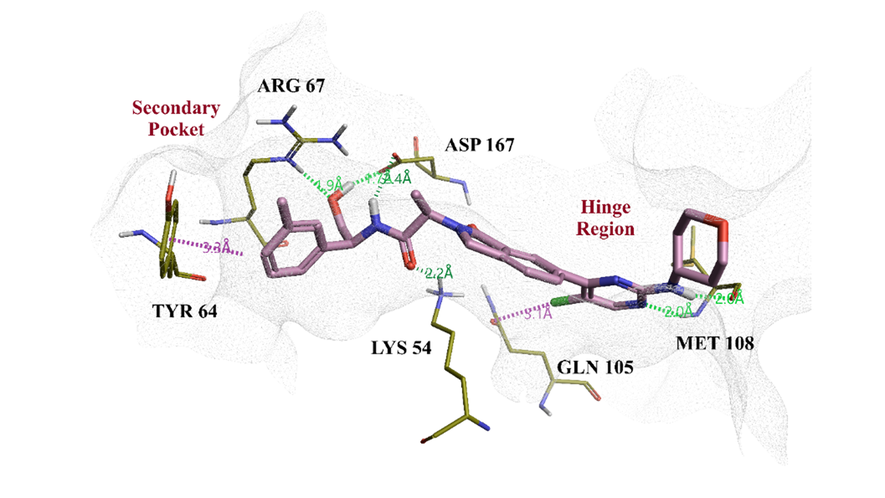
Figure 2. Compound 7 bound to the ERK2 active site (PDB code: 6G9N).
Spark suggests a list of biologically relevant replacements using Cresset’s unique molecular field technology.7 Spark replaces a specified portion in a given starter molecule with fragments derived from commercially available and literature compound collections which exhibit similar electrostatic and steric properties. The main advantage of Spark over similar tools is that it scores the whole suggested molecule instead of scoring only the replacement fragment in isolation. This product centric approach allows considering electronic and conformational effects of the replacement fragment on the rest of the molecule. Here, we will use Spark to rapidly identify alternative R-groups to the tolyl moiety (orange circle in Figure 1 – left) preserving the activity at the ERK2 target while tackling metabolic liability.
Method
The X-ray crystal structure of lead compound 7 in complex with ERK2 (PDB code: 6G9N) was used as the starting point for this case study. Spark provides wizards for setting up various experiments easily. We used the ‘Scaffold Hopping or R-group Replacement’ wizard (Figure 3 – left) to set-up the experiment.
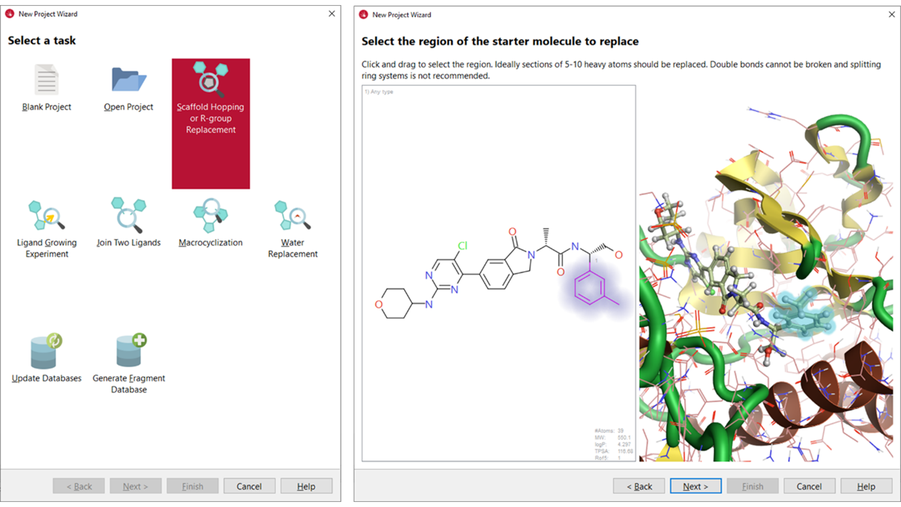
Figure 3. Left: the project wizard in Spark. Right: in the Spark wizard, the region of the starter molecule selected for replacement is highlighted by halos in the 2D and 3D window.
The PDB 6G9N was downloaded in Spark directly from the Protein Data Bank and prepared using ‘Let Spark choose the protonation state’ option. The co-crystallised ligand (compound 7) was selected as the starter molecule and all the crystallographic water molecules were deleted. The tolyl moiety was selected as the region of the starter molecule to replace (Figure 3 – right).
The calculation was run using the following settings:
- Calculation method: Accurate but Slow
- Databases to search: ChEMBL_common, Very Common and Common
These Spark databases of fragments currently contains more than 97K fragments for R-group replacement experiments which are derived from compounds reported in literature (ChEMBL database)8 and commercial compounds (VeryCommon and Common).9
Furthermore, based on the physico-chemical properties of the starter molecule (MW: 550, SlogP: 4.3, TPSA: 117, Flexibility: 8.5), the following advanced settings were used to focus the Spark search on results with druglike properties:
- Maximum number of heavy atoms: 50
-
Maximum molecular weight: 610
-
Calculated lipohilicity (SlogP) range: 3-5
-
Calculated Topological Polar Surface Area range: 40-140
Results
The R-group replacement experiment on the tolyl moiety of lead compound 7 using the Spark settings detailed above generated 1000 rank-ordered suggestions based on 50% field and 50% shape similarity, with Bio-Isostere Factor (BIF%) values ranging from 11 to 92. The BIF% is a scaled score such that the original molecule scores 100% and the score for deleting the part to be replaced is zero. Positive values indicate favorable bioisosteres, negative values reflect a replacement that can reproduce the geometry of the original molecule but is a poor mimic of the deleted moiety. The larger clusters of results include phenyl, thiophene, pyridine and furan scaffolds. Not surprisingly, the large majority of results (348) belong to the top scoring cluster of phenyl-substituted R-groups.
Phenyl-substituted R-groups
Spark results in this cluster include a number of interesting variations which can help modulate in a subtle manner the electrostatics and shape of the R-group to tackle metabolic stability, using medicinal chemistry knowledge to prioritize molecules. For example, fluorine and methoxy groups are commonly used by medicinal chemists to block a metabolic soft spot, as the carbon-fluorine bond is much more resistant to cytochrome P450 attack than the carbon-hydrogen bond. Additionally, fluorine substitution at sites adjacent to and sometimes distal to the site of metabolic attack alter the drug metabolism.6,10,11 Interestingly, the Spark results include a number of fluoro- and methoxy-substituted results, including the R-group replacements used for optimization of lead compound 7 (Table 1 and Figure 4).
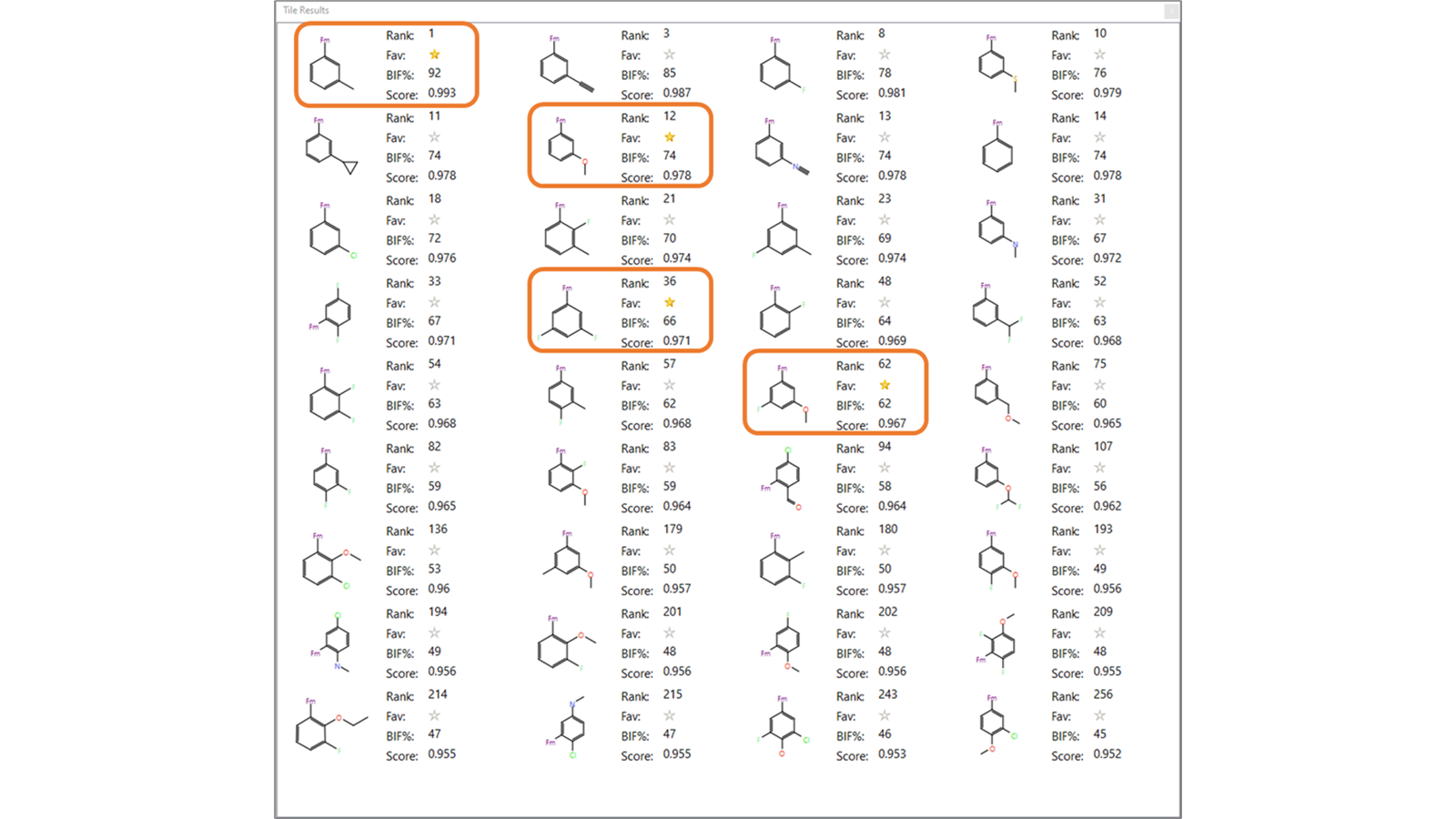
Figure 4. Tile view of some phenyl-substituted Spark results. The R-group replacements used by Astex Pharmaceuticals for lead optimization are highlighted.6
Other heterocycles
The robustness towards cytochrome P450 mediated oxidation can also be improved by replacement of an aromatic system with a more electron-deficient ring system. In particular, the replacement of a phenyl ring with a pyridyl substituent is the most common strategy to address metabolic liabilities.12 Also in this case Spark proposes a number of interesting heterocycles alternative to the tolyl moiety, including clusters of ortho-, meta-, and para-pyridines (Figure 5).
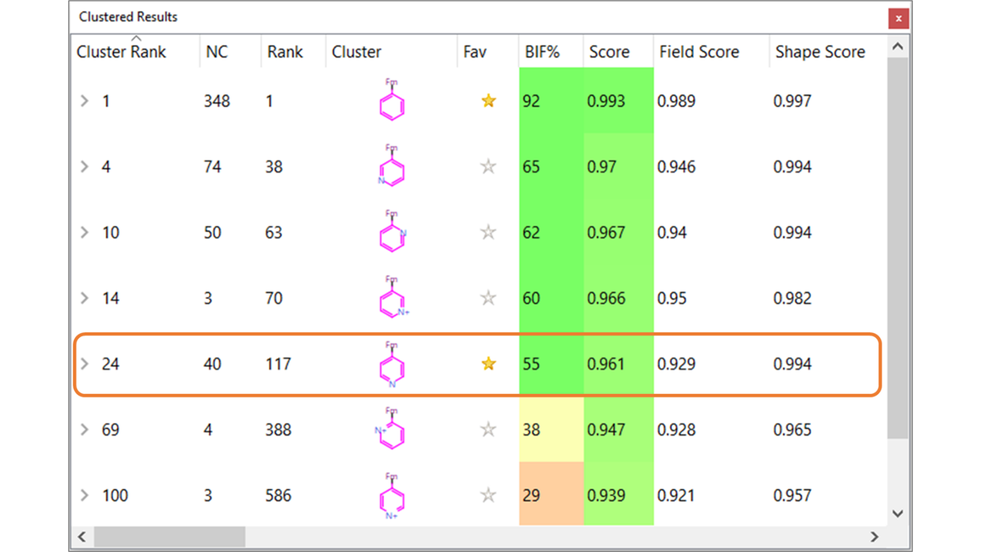
Figure 5. The top-scoring clusters in the Spark results. Note that clusters of bicyclic and five-membered heterocycles were excluded for clarity.
In particular, a cluster of 40 para-pyridine substituents, similar to those reported in the original paper6, appears with Cluster Rank 24. This cluster includes Compound 17 in Table 1, although with a lower BIF% = 46, as it is electrostatically much less similar to compound 7 than the other compounds in Table 1 (Figure 6).
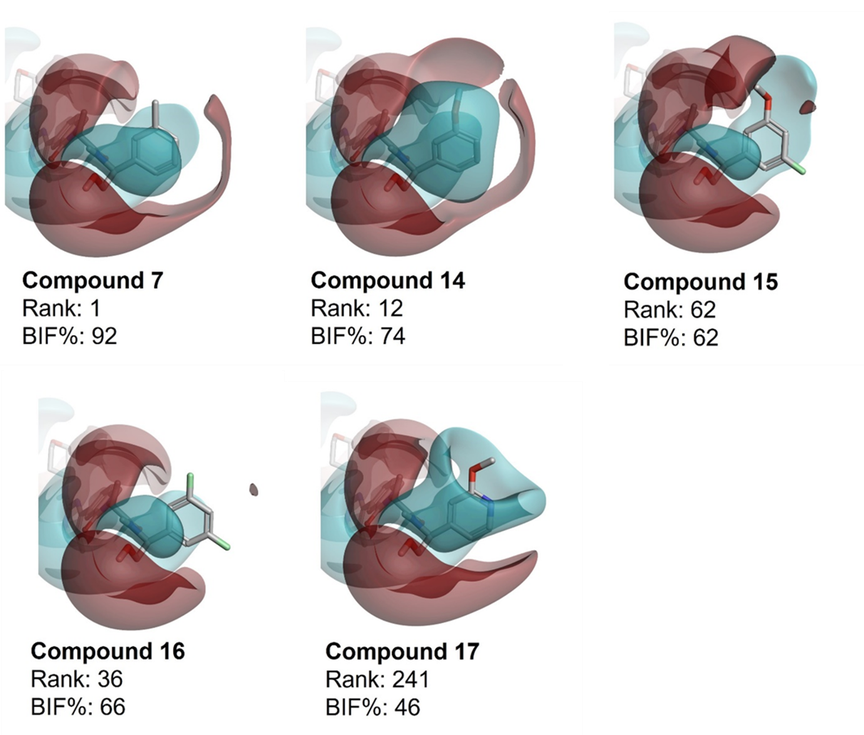
Figure 6. Comparison of positive (red) and negative (blue) electrostatic surfaces for compounds listed in Table 1. The region in which compound 17 is most dissimilar to other compounds is the negative field in front of the pyridine nitrogen atom.
Some interesting suggestions
The tolyl methyl group of lead compound 7 undergoes extensive oxidative metabolism.6 The replacement of methyl group with a cyclopropane ring can curtail cytochrome P450 mediated metabolism as proton abstraction from a cyclopropane ring would be more difficult than from a methyl group, which is needed for the oxidative metabolism.13 Among the top scoring Spark results, we can find cyclopropyl-substituted result at Rank 11 (Figure 4).
Compound 14 with anisole in place of the tolyl moiety, exhibited similar metabolic stability (Table 1). However, the replacement of anisole with difluoroanisole can improve the metabolic stability.14 The difluoroanisole-substituted result also appeared in the Spark results at Rank 107 (Figure 4). These substitutions might provide interesting alternatives to the tolyl moiety for further exploration of this lead series.
Conclusion
In this case study, Spark successfully identified the identical and very similar R-group replacements used by Astex Pharmaceuticals in the discovery of clinical candidate ASTX029.6 This example demonstrates that bioisosteric replacements with Spark in combination with medicinal chemistry knowledge could assist in addressing metabolism related liabilities in drug discovery.
References
-
Samatar A. A. et al., Nat. Rev. Drug Discov. 2014, 13(12), 928-942
-
Pearson G. et al., Endocr. Rev. 2001, 22(2), 153-183
-
Roskoski R. et al., Pharmacol. Res. 2019, 142, 151−168
-
Heightman T. D. et al., J. Med. Chem. 2018, 61(11), 4978−4992
-
Heightman T. D. et al., J. Med. Chem. 2021, 64(16), 12286-12303
-
https://www.emolecules.com/info/products-screening-compounds
-
Shah P. et al., J. Enzyme Inhib. Med. Chem. 2007, 22(5), 527-540
-
Park B. K. et al., Annu. Rev. Pharmacol. Toxicol. 2001, 41(1), 443-470
-
Lazzara P. R. et al., RSC Med. Chem. 2020, 11(1), 18-29
-
Talele T. T., J. Med. Chem. 2016, 59(19), 8712-8756
-
Xing L. et al., ChemMedChem 2015, 10(4), 715-726