With the increasing pressure to drive efficiencies in the drug discovery process, innovative approaches using computational chemistry are delivering proven results, particularly in the areas of lead identification and optimization. However, maintaining an in-house team is often a luxury and there is an increasing trend to outsource computational chemistry in order to benefit from the advantages it delivers in terms of insight into the biological activity and interactions of molecules across a range of target classes, enabling the identification of new candidates that would otherwise have been overlooked.
In silico modeling
Computational chemistry is the process of modeling chemistry in silico. It uses the principles and equations of physical and theoretical chemistry to provide solutions to problems that can be solved faster than by using traditional experiments. Key to the success of any computational chemistry experiment is the relationship between the real experiment and the computer models so that the results are trusted by the scientists in the lab.
In drug discovery, agrochemical discovery, and increasingly in flavors and fragrances, computational chemistry is used to predict the physical properties and most importantly the biological activity of compounds before they are synthesized or biologically tested.
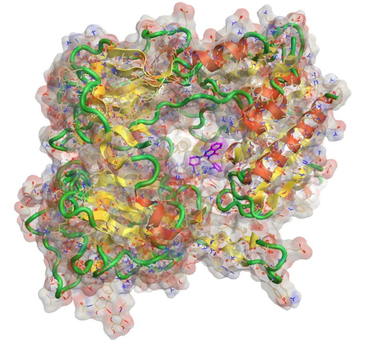
Ligand binding mode visualization showing a DPPIV ligand PDB: 2ONC. Produced using the Flare™ structure-based design platform.
Using the computer in this way saves many hours of lab time or in the case of HTS, millions of dollars in wet screening costs. Whilst the ideal of designing one molecule in silico that then becomes the drug is a long way off, new computational methods such as those invented by Cresset are making a significant impact on the discovery process.
Why outsource?
Where computational chemistry has really made a mark in practical drug discovery is in the combinatorial and high throughput screening explosion from the past two decades, in which it has become necessary to be able to process millions of molecules that are the feedstock of modern drug discovery.
There are two main factors driving the current increasing trend in outsourcing computational chemistry:
- The increasing fragmentation of the drug discovery business has brought an increase in companies that can move quickly to offer tailored research often through the use of on-demand services from consultants. At the same time, computational chemistry has become a more integral and accepted part of the drug discovery process. For example, 3D computational molecular design is now an essential part of the medicinal chemists’ toolkit.
- Big pharma companies are seeking to reduce their R&D costs as many key patents are due to expire over the coming years.
Reduce risk and increase ROI
As IP space is becoming more crowded and regulatory requirements more stringent, companies are keen to reduce risk and overheads in every way possible. Outsourcing computational chemistry can play a part in the drive to get maximum ROI on R&D spend.
In a changing drug discovery environment, where collaboration and outsourcing are key, a molecular design platform utilized by large pharmaceutical companies, biotechs, CROs and service providers can only lead to increased efficiencies in discovery.
There is a critical need for companies to efficiently convert their increasingly large base of biological targets into viable and developable drug development programmes; this now more frequently entails applying novel means of molecular hit discovery and molecular design rounds, in addition to synthesis and biological testing of those molecules.
Computational chemistry is not magic, it is science. And scientific tools need expert scientists to provide intelligent input and interpret the results.
Why work with Cresset Discovery Services?
Cresset Discovery Services‘ goal is to lower your drug discovery costs and accelerate the creation of candidates in order that serious conditions with unmet medical needs can be effectively and efficiently addressed. We help you remove obstacles, give a fresh perspective and improve your discovery performance.
Our expertize and depth of experience delivers valuable results for our customers. By outsourcing computational chemistry to us companies benefit from expert scientific knowledge, experience and methods. Outsourcing to us is very cost-effective and there is no need to go to the expense of hiring and training in-house computational chemists that may not be put to use full-time.
Contact us today for a free confidential discussion.
