An analysis of the water stability and positions in a ligand-protein complex informed the design of novel ligands for a customer target. This work led to new active chemistry that the customer went on to patent.
A Cresset Discovery Services customer had identified a novel target with a natural ligand and were looking for new chemistry that would be active at the target site. Our scientists carried out an initial project to learn more about the protein-ligand system. The Cresset field approach, used to analyse the structure and interactions, gave the customer valuable insights into the active features of the ligand.
The customer used this information to develop analogous synthetic compounds and example molecules. They asked us to work with them again to computationally align the example molecules and prioritize them for synthesis.
We carried out an initial alignment and then modeled the system in detail. It appeared that part of the molecule that was important for the interaction was not making any contact with the protein.
The PDB had some crystal structures of related proteins, but not of the target of interest. We studied the available protein data to learn as much as possible about the binding pocket, paying particular attention to the positions and stability of the water molecules. This led to us putting forward the hypothesis that an important part of the ligand interaction included the stabilization of water.
Based on this hypothesis we prioritized the molecules that bridged the observed gap between the natural ligand and the target while also stabilizing the free waters.
Water analysis was carried out by manually superimposing multiple crystal structures, viewing the crystallographic waters that clustered together, and mapping on their temperature factors. This process allowed us to determine the importance of each water molecule in the solvation sphere around the ligand and protein pocket. With the advent of the new 3D-RISM method in Flare a similar computational work-flow can be accessed which is far more efficient for this type of analysis. This is a more systematic approach which enables us to calculate the position and stability of all water molecules around a proposed ligand in a binding pocket. Moreover, as it does this without the need for any crystallographic water data, this is far more useful as well as convenient. Ultimately, this data can be used to assess or compare ligands in terms of how well they might stabilize essential water.
Based on our equivalent ‘hands-on’ analysis, we worked with the customer to choose the best candidates for synthesis. These newly-designed ligands resulted in new active chemistry for the customer that was valuable enough for them to patent.
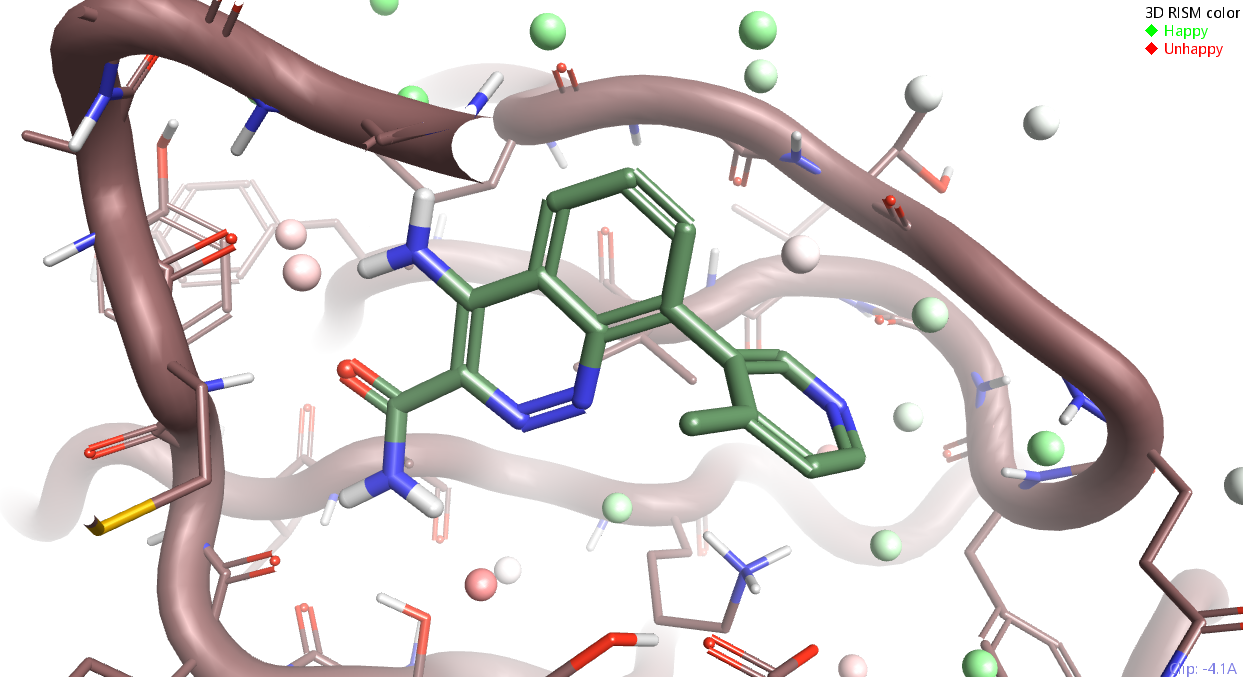
The position and energetics of water molecules in and around the active site is of crucial importance when designing novel ligands. Knowing which water molecules are energetically favorable can give valuable insights into the best positions for ligand molecules. 3D-RISM analysis is one of the methods available in Flare for structure-based drug design.
