Virtual Screening Strategies for Identifying Novel Chemotypes
In their recent article in Journal of Medicinal Chemistry, Stuart Lang and Martin Slater review in silico approaches for identifying ...
News
Cresset’s software is underpinned by a proprietary force field which was designed specifically to provide a more accurate description of the electrostatic fields around molecules. This science is well established in the pharmaceutical industry where Cresset has a global customer base. Our molecular fields intrinsically relate more closely to the properties involved in molecular recognition events operating at the level of the biological target, be that a protein receptor or enzyme.
Flavor and fragrance molecules, similarly, are also required to interact with specific protein targets to exert their effects and thus our field based techniques are theoretically equally applicable. Indeed, we are already working with a number of flavor and fragrance companies in our capacity as a provider of molecular modeling consulting services.
The main biological targets of fragrances are thought to be the human olfactory receptors which belong to the wider, therapeutically important, GPCR family. Flavors are more diverse and complex in their interactions, with both GPCR’s and ion channel targets known, but also components of taste perception include concomitant triggering of olfactory receptors.
Clearly, there are many differences between flavor and fragrance molecules compared with their drug counterparts. Pharmacological effects cover a wide spectrum of therapeutically relevant biological targets, and also require physiochemical properties in keeping with their use, e.g. oral, CNS, inhaled or topical applications.
Fragrance molecules need to be non-toxic and volatile and as such are restricted by tight chemical and physical property considerations. In addition, the mode of action of fragrance molecules is generally accepted to be through GPCR agonism and thus the spectrum of size, shape and mwt is accordingly further narrowed (Figures A and B). It is an incredible observation that the huge spectrum of fragrance perception is dictated by the precise 3D distribution of typically between only 2 and 20 atoms of only five elements CHNO and S. Possibly, unsurprisingly, this translates into very compact electrostatic, shape and chirality profile for each of these special molecules.
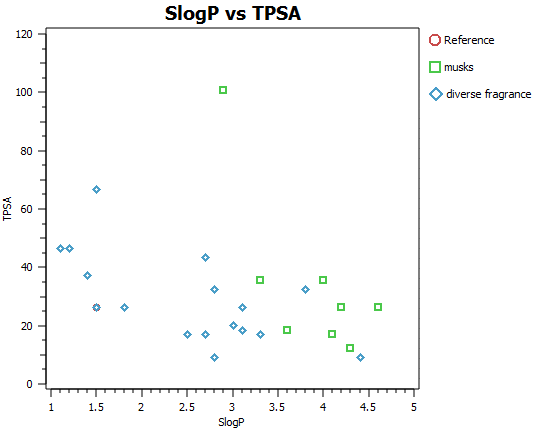
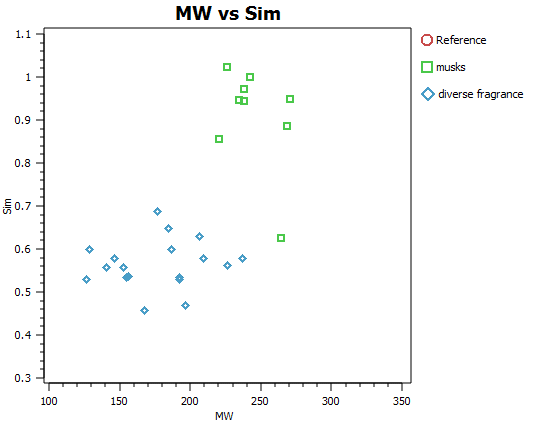
In order to illustrate the usefulness of Cresset’s electrostatic descriptions for fragrance molecule space, a small set of 27 diverse fragrance molecules (Kraft 2000, Angew Chem. review and the Leffingwell and Associates website) were built and analysed in our ligand focused molecular modelling software suite Forge.
Alignments were made and all set, to fix the centre of gravity and the overall dipole, relative to an arbitrary starting point ‘Coumarin’. Similarity scores were then re-calculated relative to ‘Ambrette’, which is one of the largest example ligands, to ensure sensible (non-negative) similarity values. Among the set an extended group of Musks were included and analysed. These alignments were refined by hand – and are representative of the workflow we would typically employ for developing each fragrance grouping – given more time.
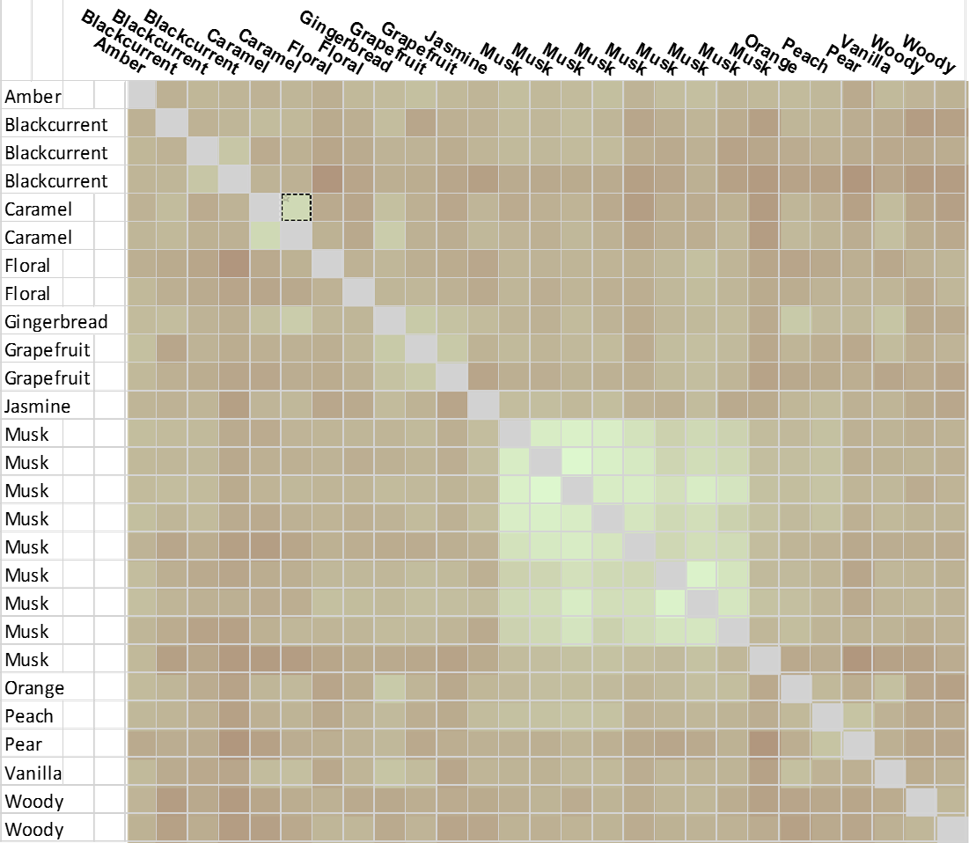
In the analysis, the effect of picking a single reference is not critical since each molecule was scored relative to every other, using Forge’s Activity Miner module, to generate an all-by-all matrix (Figure C). This simple exercise is limited only by the low numbers involved in the analysis, yet still provides a very intuitive picture. From the matrix, brighter ‘green’ corresponds to higher similarity, darker brown – low similarity.
There is a significant cluster around the musks, but also other hot spots of high similarity. Two example ‘caramel’ fragrance molecules (highlighted as a pair in the matrix (black box) are shown as 3D aligned molecules (Figure D).
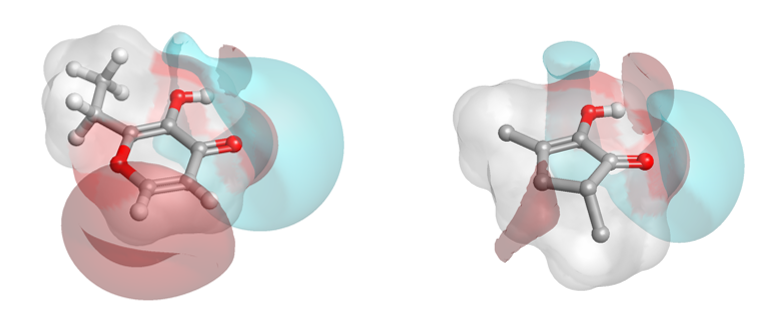
In stark contrast, the purely chemical structure driven analysis (using EFCP4 2D similarity, Figure E) shows mostly low similarity. Unsurprisingly, as the relationship of the musk SAR here would likely defeat most 2D similarity methods, the structurally diverse Musk clusters disappear and the caramels blend into the noise.
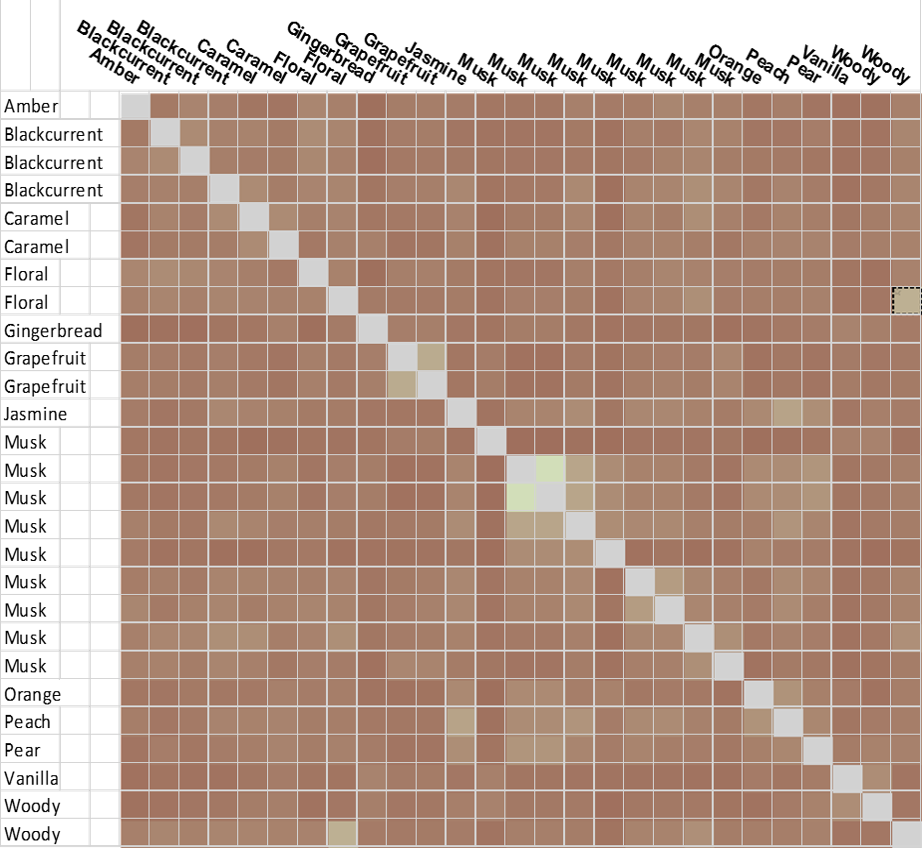

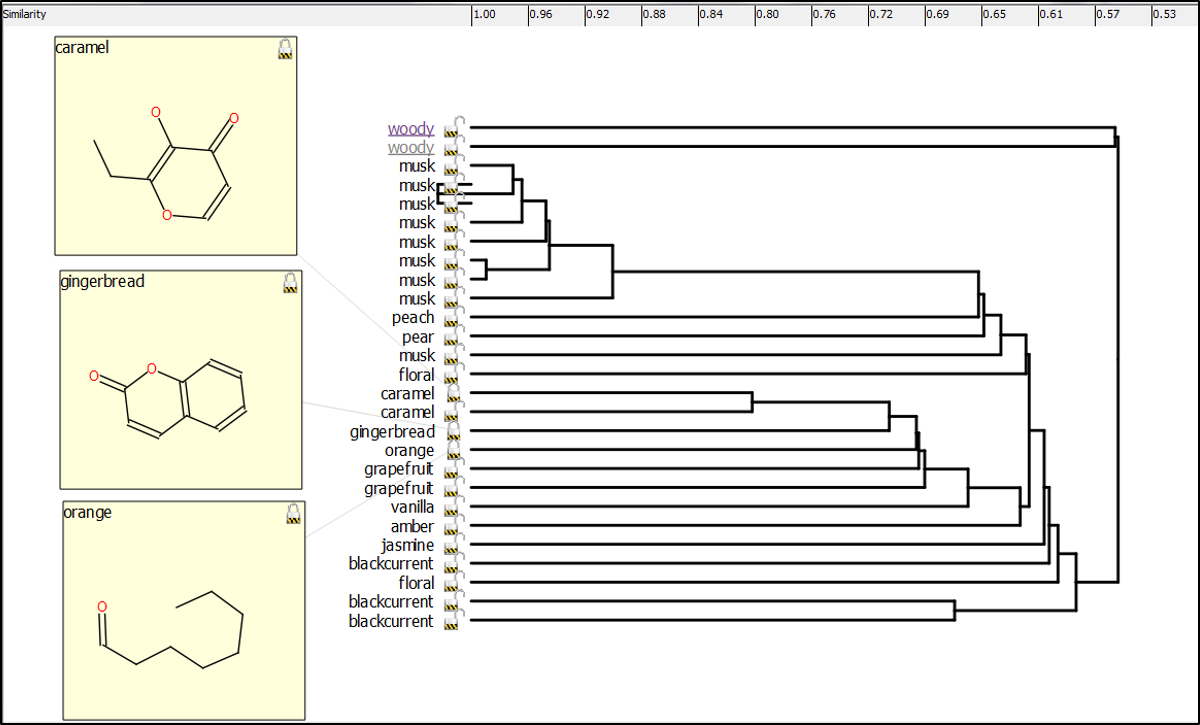
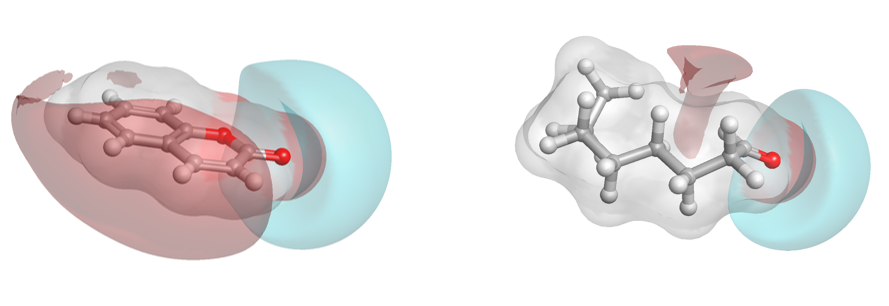
Forge was also used to generate the hierarchical clustering (Figure F) which again provides an intuitive view of the fragrance set. Fascinatingly, gingerbread, caramel and orange fragrance examples, which would not jar from a gustatory sense, show a similarity suggesting that their corresponding targets may be more similar to each other than others (3D aligned structures in Figure G).
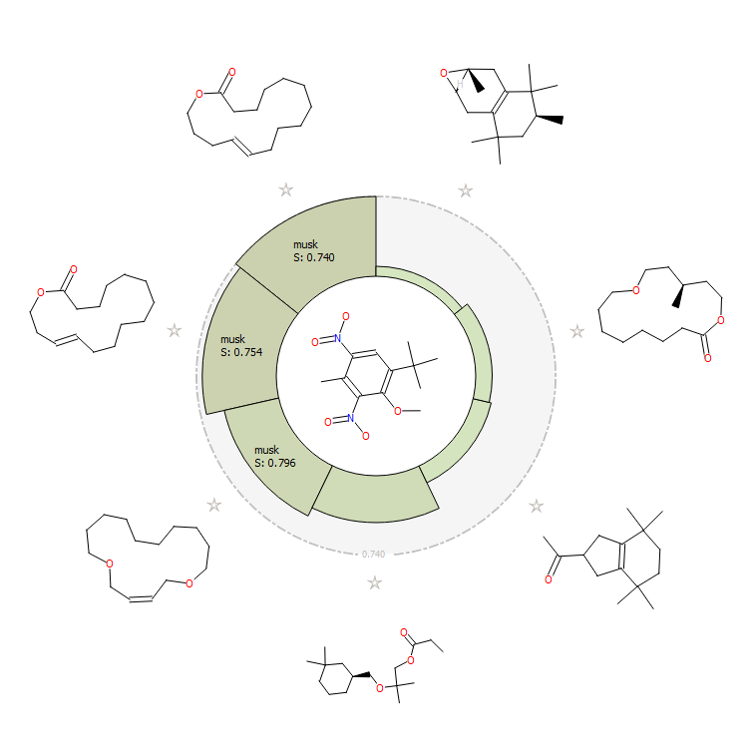
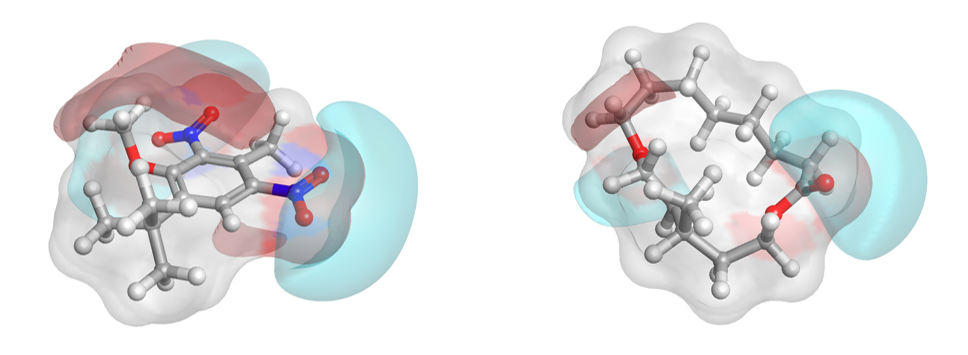
Additionally, the activity view from Forge (Figure H) shows visually how nicely the nitro musk relates, in a field similarity sense, to the chemically distinct acyclic, bicyclic and macrocyclic musks. Two chemically distinct musks molecules are shown in 3D (Figure I).
In conclusion, this analysis demonstrates the power of Cresset’s technology for usefully analysing bioactive molecules. The way in which we can do this gets to the essence of the way the molecules interact with their targets – independently from the chemical architecture.
Field based models of active flavor and fragrance ingredients could potentially provide novel chemotypes for this industry much in the way that it has provided new hits and leads for the pharmaceutical industry.
See more about Forge’s Activity Miner module or try a free demo of Forge. Alternatively contact us for a free confidential discussion.