The adaptability of pyFlare to access advanced data visualization and calculation functionalities
We showcase some examples within a drug discovery context where the pyFlare environment has been further expanded with advanced data ...
News
We are delighted to announce the release of Flare beta 2. This version has many enhancements suggested by users as part of the on-going beta test program and is available for evaluation from your account manager. This final round of beta testing will focus on fine tuning the operation of Flare – perfecting keyboard shortcuts, adding more quick access items and polishing dialogue boxes in the run up to launch. So you have an application that meets your needs, we are interested in hearing about where you think the application can be improved.
Since the first beta test we have made a number of improvements both in response to your feedback and from our own experience. One of the most significant changes is an overhaul of the relationship between ligands in the ligand table and their parent protein. In Flare beta 2, each ligand has a parent protein that is set automatically and can be manually adjusted by simply double clicking the table cell. This enables ligands to be grouped together by chemistry, source, or parent protein making full use of the ‘Molecule roles’ feature.
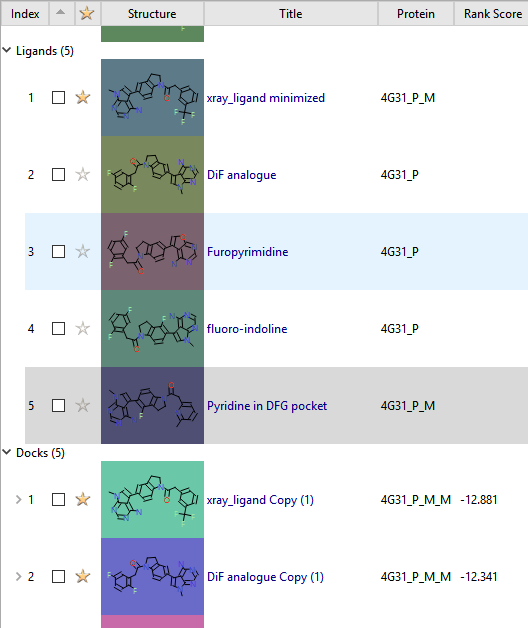
Molecules in two roles within the ligand table with their Title, associated Protein, and Rank Score from docking.
All the calculation dialogues have been significantly improved to enable parallel processing and more visual feedback on the extent of the calculations. Now, whenever you setup a calculation the 3D window will display relevant calculation boxes, from the size of an active site in a docking experiment to the clipping boxes for surface generation.
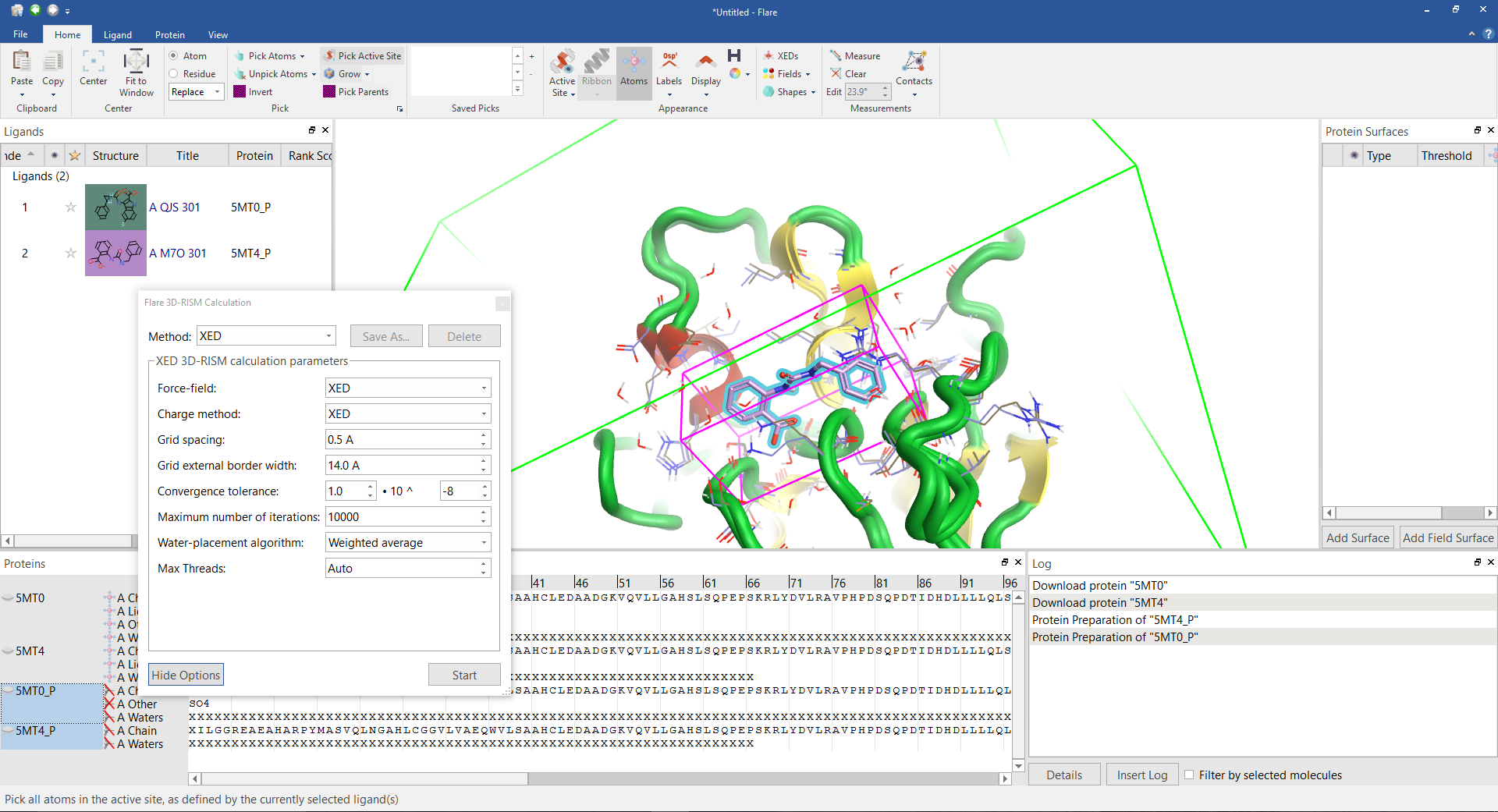
A 3D RISM calculation in preparation showing the cube in which the RISM waters will be placed (magenta) and the hydration shell that surrounds the calculation (green).
The contact detection and display algortithm have been overhauled to give significantly greater performance and to show only the contacts that you are interested in. Flare now gives control over the display of individual interaction types, whether to include waters, and the inclusion of intramolecular interactions (such as H-bonds within a protein).
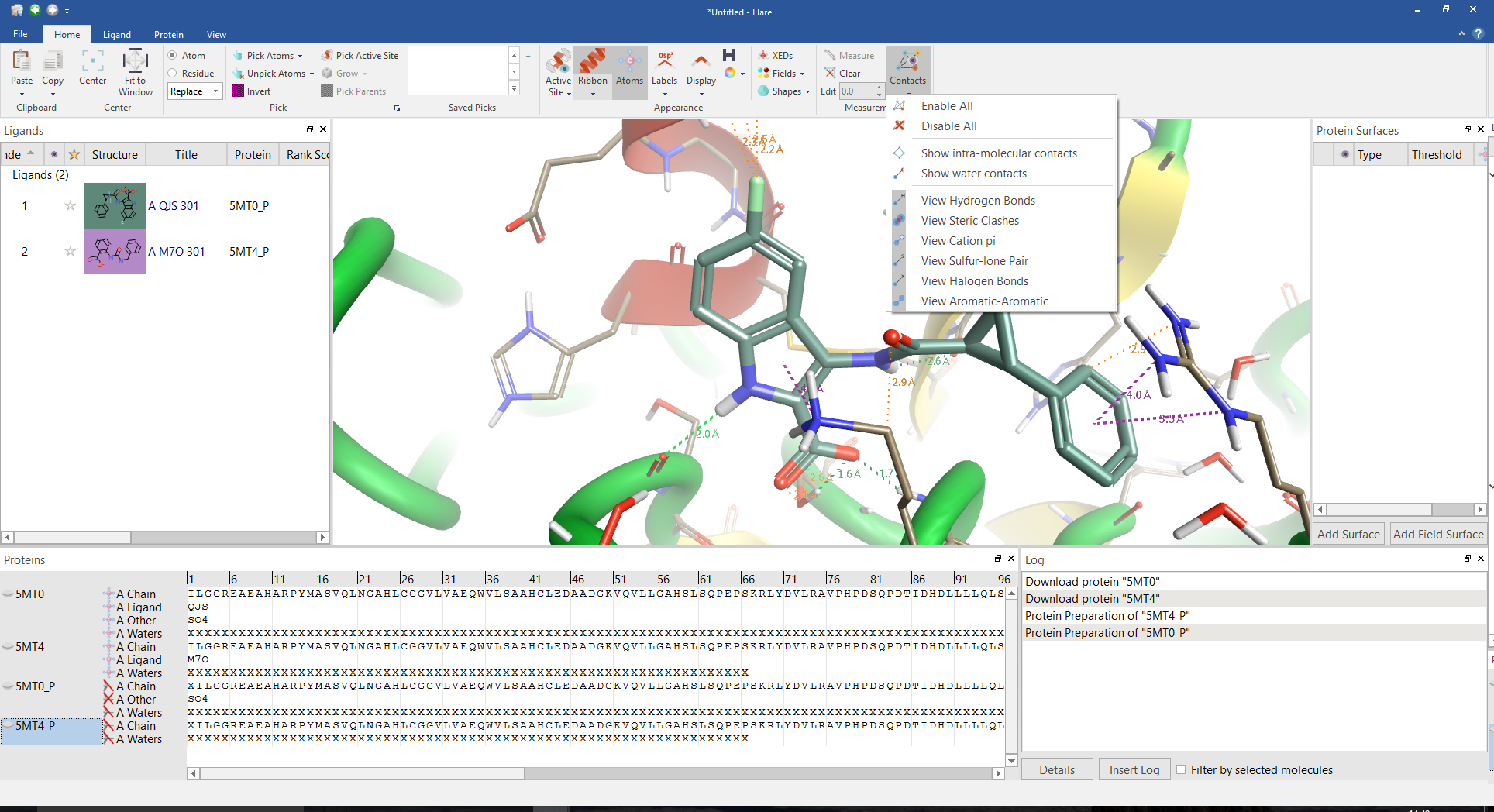
Interactions for the ligand from PDB 5MTO.
Finally, significant work has been put into job parallelization, particularly for WaterSwap. Here we have rewritten our unique Engine Broker that enables client machines (be they Windows®, MacOS® or Linux®) to use remote or cloud based compute resources to super-power their calculations. Using the Cresset Engine Broker (CEB) starting a cloud based calculation could not be simpler:
The new CEB has a completely different architecture such that it now handles all communication. This is particularly useful when running on the cloud or other situations where the client machine knows nothing of, or cannot communicate with, the individual calculation nodes of the cluster. For WaterSwap we have modified the algorithm to make full use of cloud resources where the perfect situation is to have an infinitely wide calculation that completes in seconds. For a monte-carlo based simulation there is a limit to how wide we can make the calculation but we do not have to limit ourselves to a single process either. In Flare Beta 2 we have enabled an option to split the WaterSwap job into parallel chunks that utilize the highly parallel nature of cloud resources to run the same simulation upto 4 times faster.
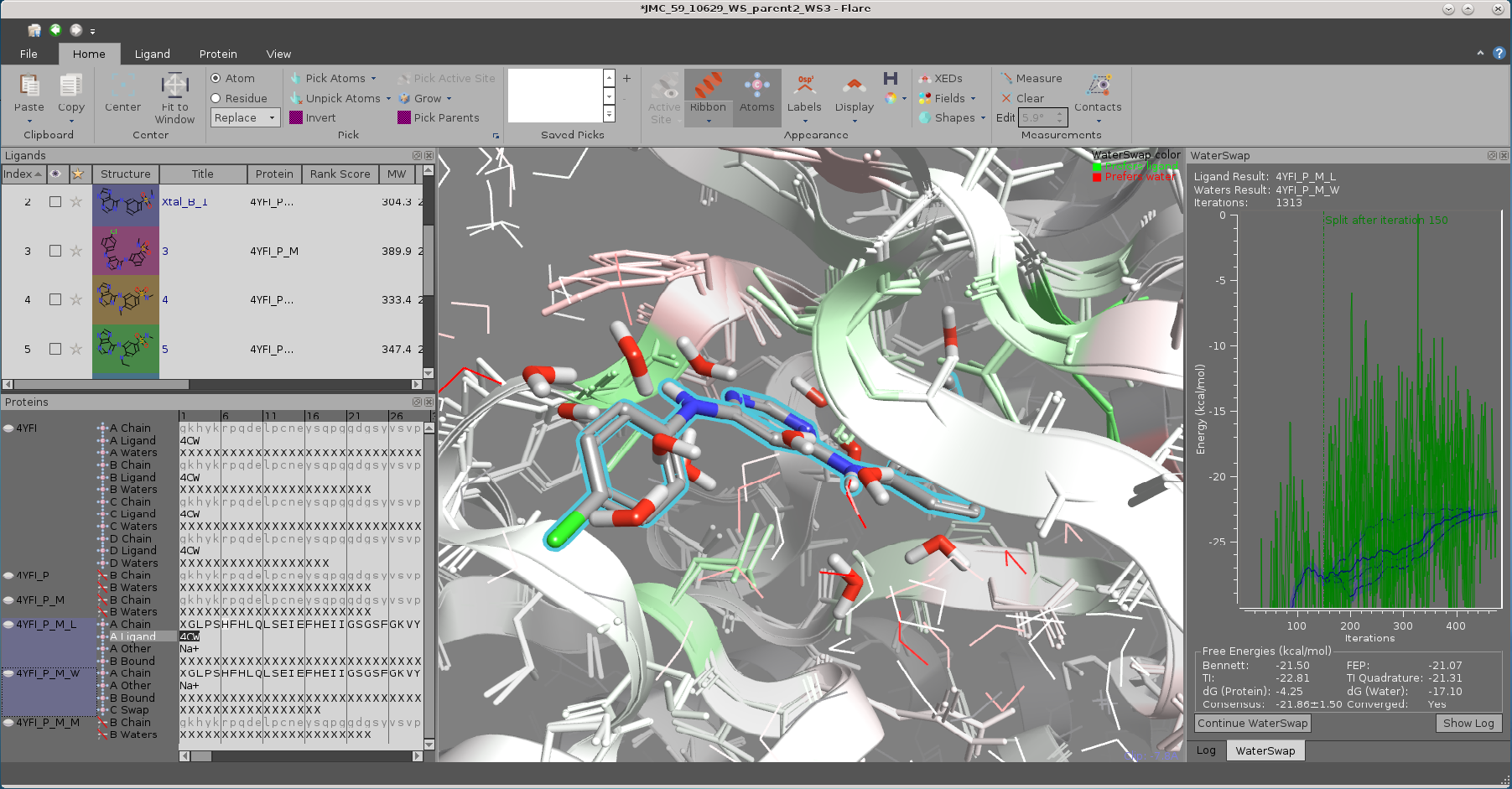
WaterSwap result for a ligand bound to TNNI3K (PDB 4YFI) showing both the ligand bound and water bound protein results from a WaterSwap experiment.
Interested in Flare? Contact your account manager to join the Flare beta 2 program and gain early access to this cutting edge structure-based design method with intuitive GUI.