Improving PROTAC properties via single-point changes to linkers
We explore how computational methods can be applied to proteolysis targeting chimera (PROTAC) design, to effectively tackle some of the ...
News
Scaffold hopping and R-group replacement remain central tasks in medicinal chemistry for generating and protecting intellectual property. Spark is a bioisostere replacement tool (available as a desktop software application) for rapidly generating reasonable yet novel scaffold and R-group replacements using Cresset’s molecular field points.
Cresset’s field technology condenses the molecular fields down to a set of points around the molecule, termed ‘field points’. Field points are the local extrema of the electrostatic, van der Waals and hydrophobic potentials of the molecule.
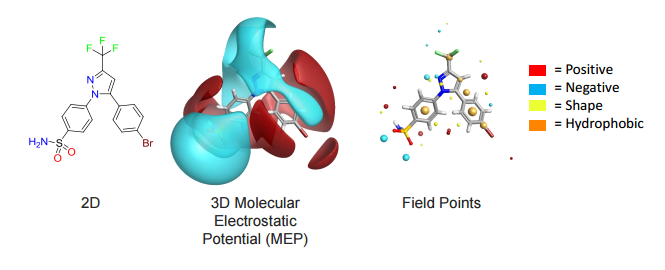
The Spark approach uses a database of molecule fragments, or available reagents, to suggest replacements that maintain the shape and electrostatic character of a known active molecule. The user identifies the region of a known active
molecule that they wish to replace, and this piece is removed.
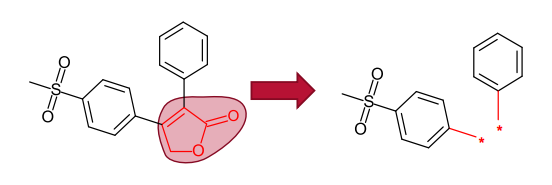
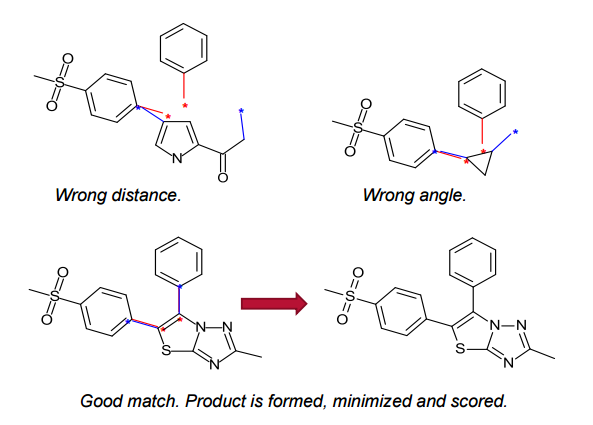
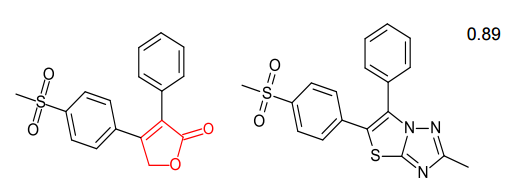
Spark generates bioisosteres from databases of fragments derived from:
In this case study we investigate which of the fragment sources available in Spark is the best source of inspiration.
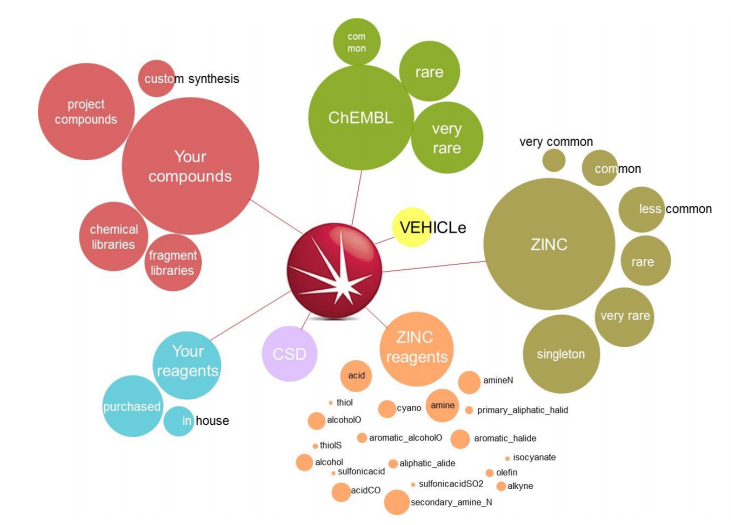
The ChEMBL ‘common’, ‘rare’ and ‘very rare’, ZINC ‘very common’, ‘common’, ‘less common’, ‘rare’, ‘very rare’, ‘singleton’ and the VEHICLe fragment databases were searched using ‘Accurate But Slow’ calculation settings. Compounds with piperazine scaffolds were filtered out as these are very well known in the literature.
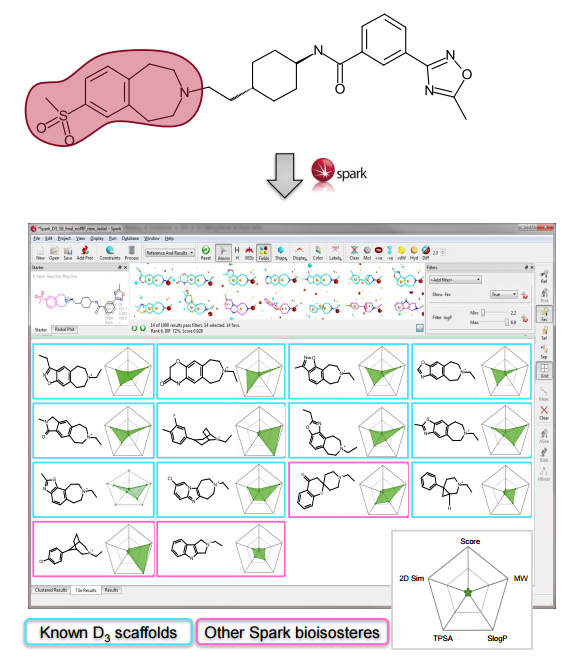
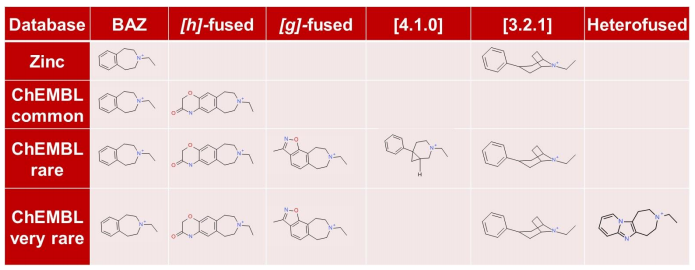
An analysis of the chemical diversity of the known D3 scaffolds retrieved from each database clearly shows that the less common fragments derived from the literature database are a precious source of potentially
useful chemical diversity. Note that these less common fragments may be associated with more complex and less documented synthetic routes.
The ChEMBL and VEHICLe fragment databases were searched using ‘Accurate But Slow’ calculation settings. The protein structure for 1UDT was used as an excluded volume, constraining the field points associated with the interaction with glutamine (Gln817) in the 1UDT protein.
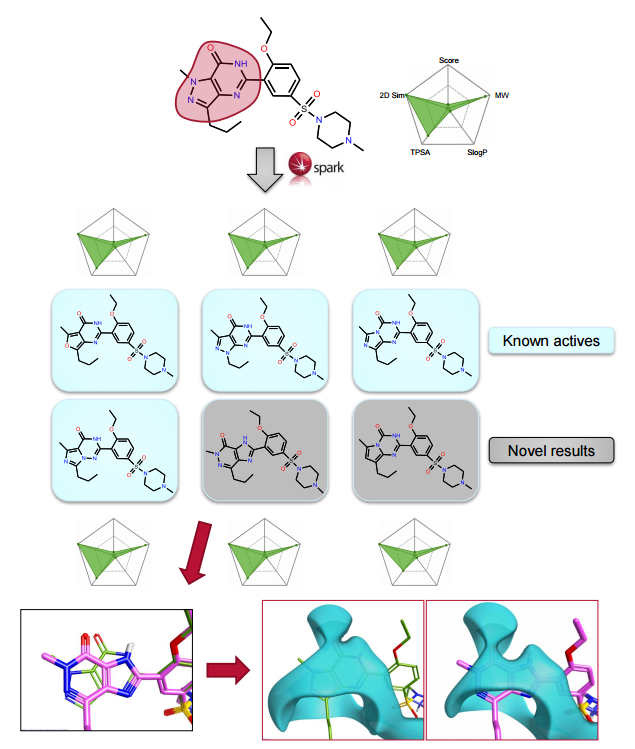
Spark provides both known active scaffolds and novel solutions that represent opportunities for scaffold hopping and R-group replacement.
The nature of the experiment appears to dictate the best source of fragments. It is therefore important to have a wide range of fragment sources to choose from for each experiment, to provide a balance between novelty and synthetic accessibility.
The creation of fragment databases from proprietary collections of compounds can be a powerful way of increasing the chemical diversity available to Spark.