Improving PROTAC properties via single-point changes to linkers
We explore how computational methods can be applied to proteolysis targeting chimera (PROTAC) design, to effectively tackle some of the ...
News
Balancing novelty, activity, physicochemical properties and IP position is at the heart of the hit-to-lead and lead optimization processes. Generating many small changes to a structure rarely moves the project forward significantly while large changes can cause a significant loss in activity. The challenge is often to find a non-trivial change that progresses the project towards the multi-parameter optimization goal without jeopardizing activity.
Bioisosterism has proved a popular way to generate new scaffolds in drug discovery. In this poster we explore the application to R-groups and demonstrate that groups which are bioisosteric in shape and electrostatic space provide an excellent range of lead optimization
opportunities. However, in hit-to-lead and lead optimization projects, there is rarely time to scope out new synthetic routes for the introduction of each R-group.
Linking in silico generated ideas for R-group replacements with available reagents and accessible chemistry is key to using the novel results available through shape and electrostatic bioisosterism. R-group libraries that are united by specific chemistry provide a systematic way to rapidly exploit a particular chemical reaction to generate novel chemical matter during the lead optimization process.
Cresset’s field technology1 represents molecules using electrostatic and shape properties enabling the comparison of molecules across chemical series.
Spark searches databases of fragments for replacements for part of the starting molecule. All fragments which have the required geometry are formed into a product molecule which is energetically minimized. Only as a
product molecule is the replacement assessed for electrostatic and shape similarity to the starting molecule. This enables the electrostatic and shape properties of the fragment influence those of the retained portions of the molecule and vice-versa
Spark generates bioisosteres from databases of fragments derived from:
We present a simple technique to rapidly generate and use databases of available reagents. It is applied to a collection of compounds to generate a searchable database of bioisosteric replacement groups to boost novelty in design, while simultaneously balancing physicochemical property and synthesis considerations. A simple set of rules classify the R-group collections by specific chemistry making selection of the appropriate database facile. Other secondary data is included in the substituent record, reflecting its source compound in an inventory system or vendor catalogue for ease of access.
In this example Spark was used to look for reagents that were bioisoteric with a pyridyl-water complex. Smith et al.<sup>2</sup> showed that the replacement of the 4-methylpyridin-3-yl in PDB:4ZLZ with small bicyclic heterocycles improved potency. The new hetrocycles displace the water molecule and make direct H-bond interactions with the P-loop.
We sought to test if Spark could suggest known and reasonable alternative replacements for the pyridyl water complex. We weighted the scoring towards electrostatics and specified the H-bond interactions of the water molecule as required. Fragments were selected from a database of 41K aromatic halides to replicate the boronic acid chemistry used in the original publication.
In this experiment we wished to demonstrate the use of Spark for providing novel amines for published D3 antagonists. However, searching for new amines from the secondary amine reagent database failed to provide novel
R-groups. Reasoning that the known active R-groups were highly functionalized and therefore that commercially available amines represented a limited source of inspiration the search was expanded to encompass all supplied fragment databases. Compounds with piperazine scaffolds were filtered out as these are well known in the literature and diluted the results.3
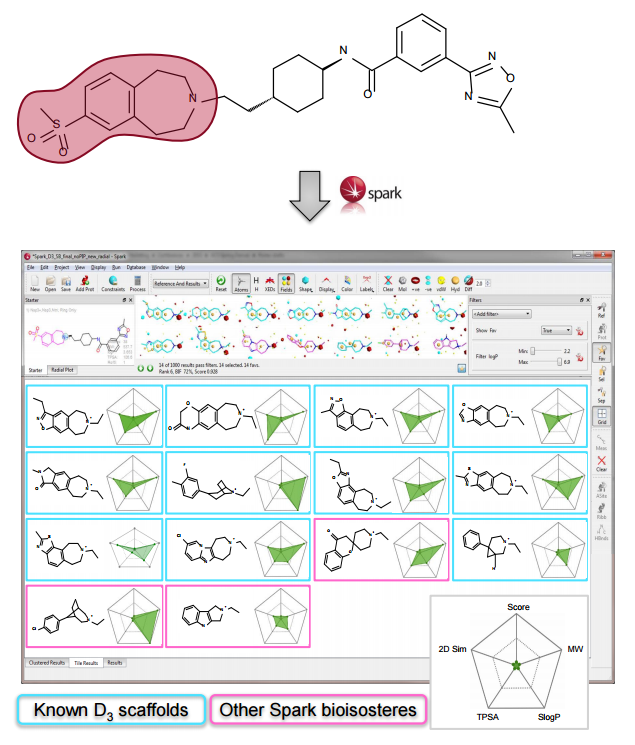
Spark provides both known and novel active scaffolds that suggest opportunities for scaffold hopping and R-group replacement. Combining this power with the use of ‘chemistry-aware’ reagent fragment databases allows for exploitation of specific chemistries in the laboratory. Accessing fragments for potential substitutions from both literature and commercial sources represents a way to identify potentially novel chemistry and diversity. Furthermore, the creation of fragment databases from proprietary collections of compounds can be a powerful way of increasing the chemical diversity available.
1. J. Chem. Inf. Mod. 2006,46, 665-676
2. J. Med. Chem. 2015, 58, 5437−5444; Nature 2003, 423 (6937), 356−361; J. Am. Chem. Soc. 1989, 111 (1), 314−321
3. J. Med. Chem. 2007, 50, 5076-5089; Bioorg. Med. Chem. Lett. 2008, 18, 901– 907; Bioorg. Med. Chem. Lett. 2008, 18, 908–912; J. Med. Chem. 2010, 53, 7129–7139