The adaptability of pyFlare to access advanced data visualization and calculation functionalities
We showcase some examples within a drug discovery context where the pyFlare environment has been further expanded with advanced data ...
News
Quantum mechanics (QM) minimization and single-point energy calculations offer a highly accurate approach to compute the minimized structures and energies of molecules. Accurate QM calculations in Flare™ are based on the Psi41 implementation, and offer the choice between Hartree-Fock (HF), Møller–Plesset perturbation theory of the second order (MP2) and Density-Functional Theory (DFT) for structure optimization and computation of molecule energetics, with DFT representing a popular choice in the field of quantum chemical structure prediction because of its exceptional accuracy.
However, DFT minimization and single-point energy calculations typically scale O(N3) where N is the number of electrons in the simulation. This means that, even when keeping the calculation relatively simple by using the very popular B3LYP hybrid exchange-correlation functional and a minimal 3-21G basis set, the computational complexity of the DFT minimization and single-point energy calculations very rapidly increases with the size of the ligand. This translates to much longer computer times as the number of atoms and, subsequently, the ligand molecular weight increases (Figure 1).
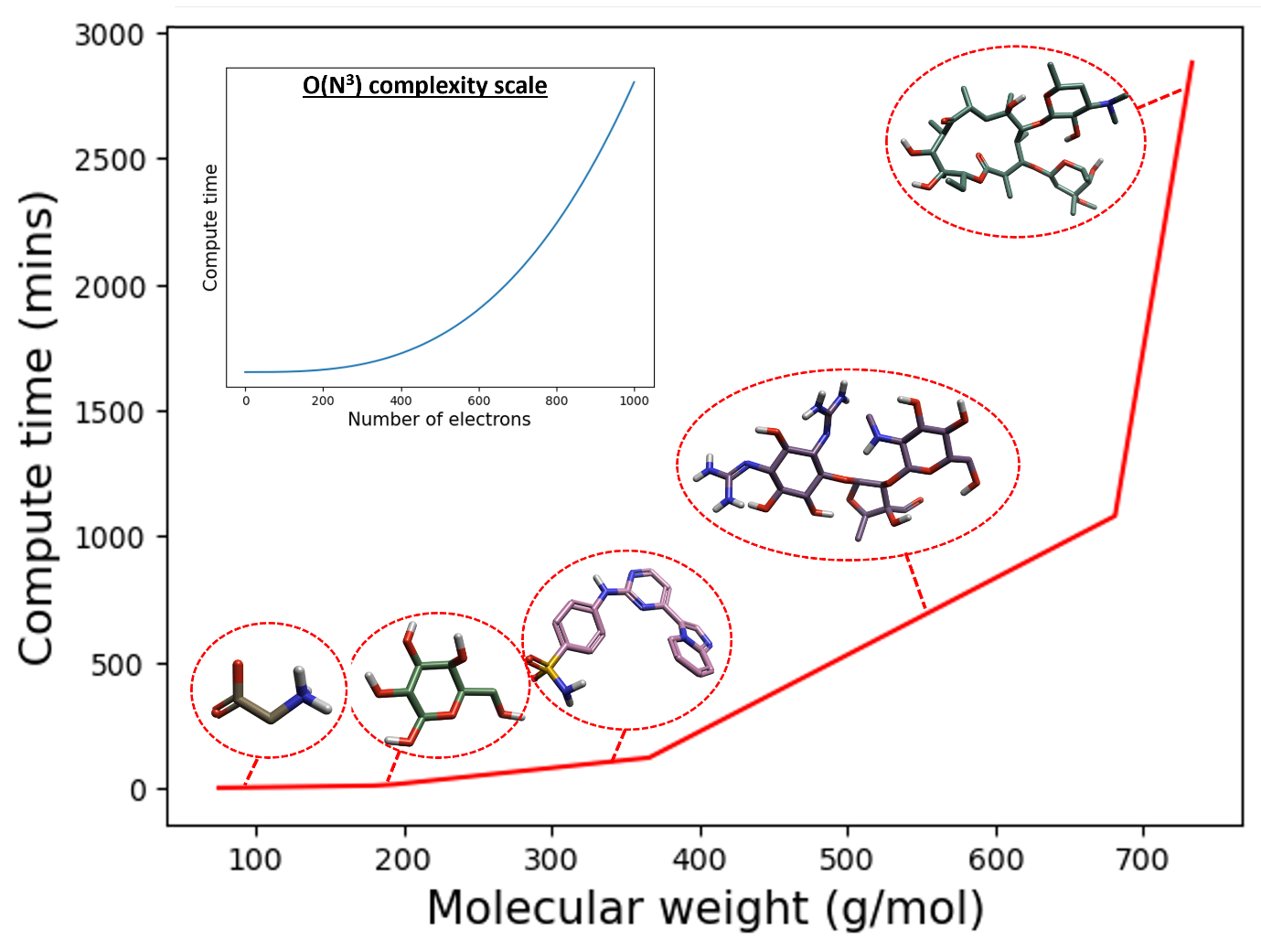
Figure 1: DFT minimization and single-point energy compute time (mins) as a function of increasing molecular weight (g/mol) for example biomolecules. From left to right: glycine, glucose, 4-[(4-imidazo[1,2-a]pyridin-3-ylpyrimidin-2-yl)amino]benzenesulfonamide, streptomycin, and erythromycin. The inset displays a generalized O(N3) relationship for comparison. The DFT compute times were calculated for an 8-core CPU.
From the left of Figure 1, it is easy to see that DFT minimization and single-point energy computations for small molecules are, indeed, rather computationally tractable. However, when moving to the right of Figure 1, and we enter the realm of macrocycles and macromolecular structures, these calculations become infeasible without access to high-performance computing (HPC) facilities. As such, for large molecules, a more computationally tractable approach to compute minimized structures and energetics is required.
In Flare, it is possible to use GFN2-xTB as an alternative theoretical method to structural minimization and computation of ligand energetics. GFN2-xTB is a semiempirical tight-binding method, designed for the fast computation of molecular energies of systems encompassing tens or hundreds of atoms.2,3 GFN2-xTB is designed to bridge the gap between QM and molecular mechanics (MM) forcefield methods, essentially enabling the modeler to compute molecular structures and energetics with accuracy closer to the former and the computational efficiency of the latter.
The GFN2-xTB calculation method is accessible from the ‘QM’ button under the ‘3D Pose’ tab in Flare (Figure 2).
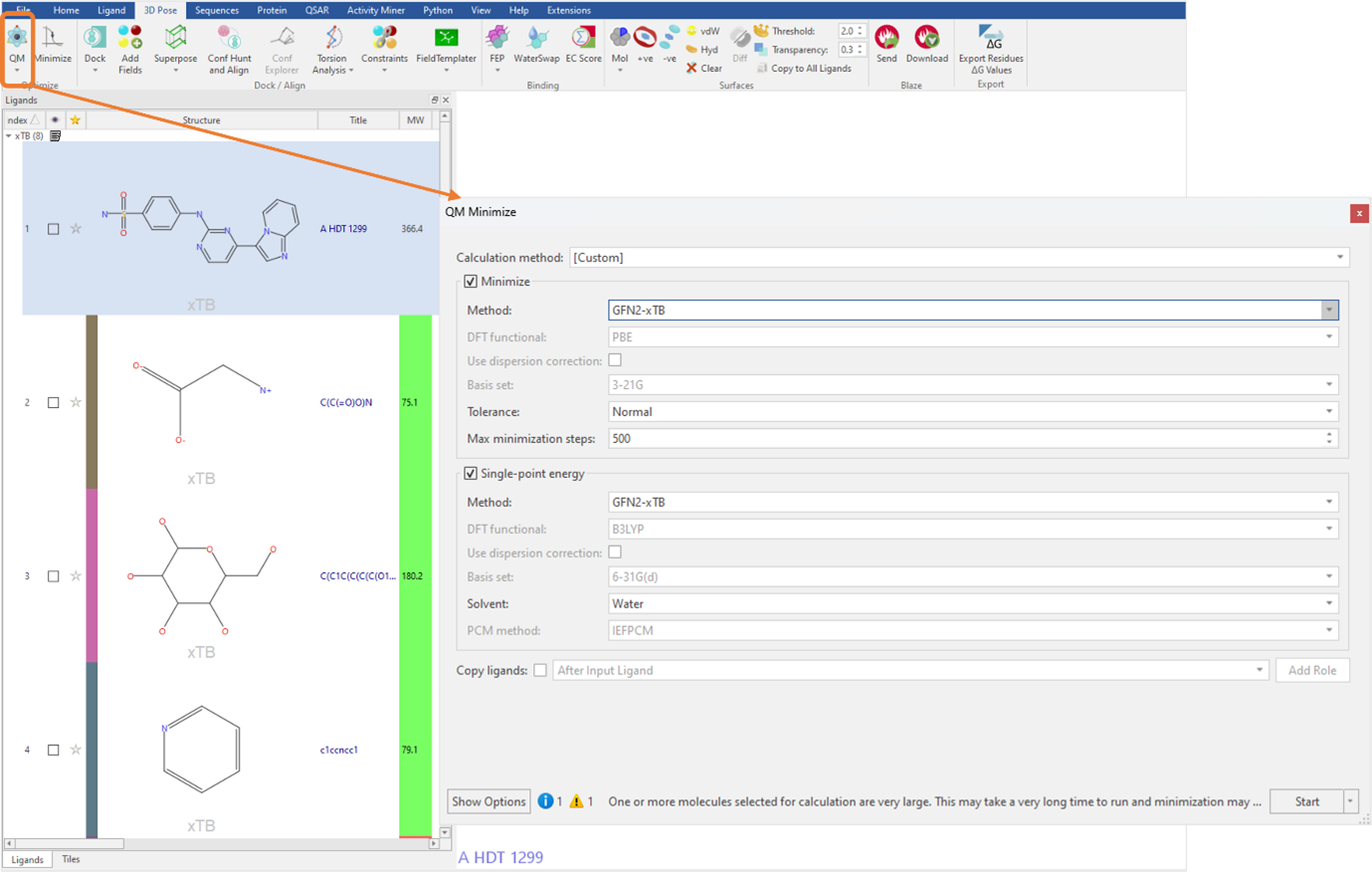
Figure 2: GFN2-xTB minimization and single-point energy calculations in Flare.
We have the option to carry out both minimization and single-point energy calculation using GFN2-xTB. Alternatively, for increased accuracy, we can construct a hybrid workflow to carry out the minimization at GFN2-xTB level of theory followed by an energy calculation at B3LYP/6-31G(d) level of theory (Figure 3). In this workflow, the most compute-intensive portion of the calculation, the minimization, is computed at a more efficient GFN2-xTB level of theory with the less compute-intensive portion of the calculation, the single-point energy calculation on the minimized structure, performed at the more accurate DFT level of theory.
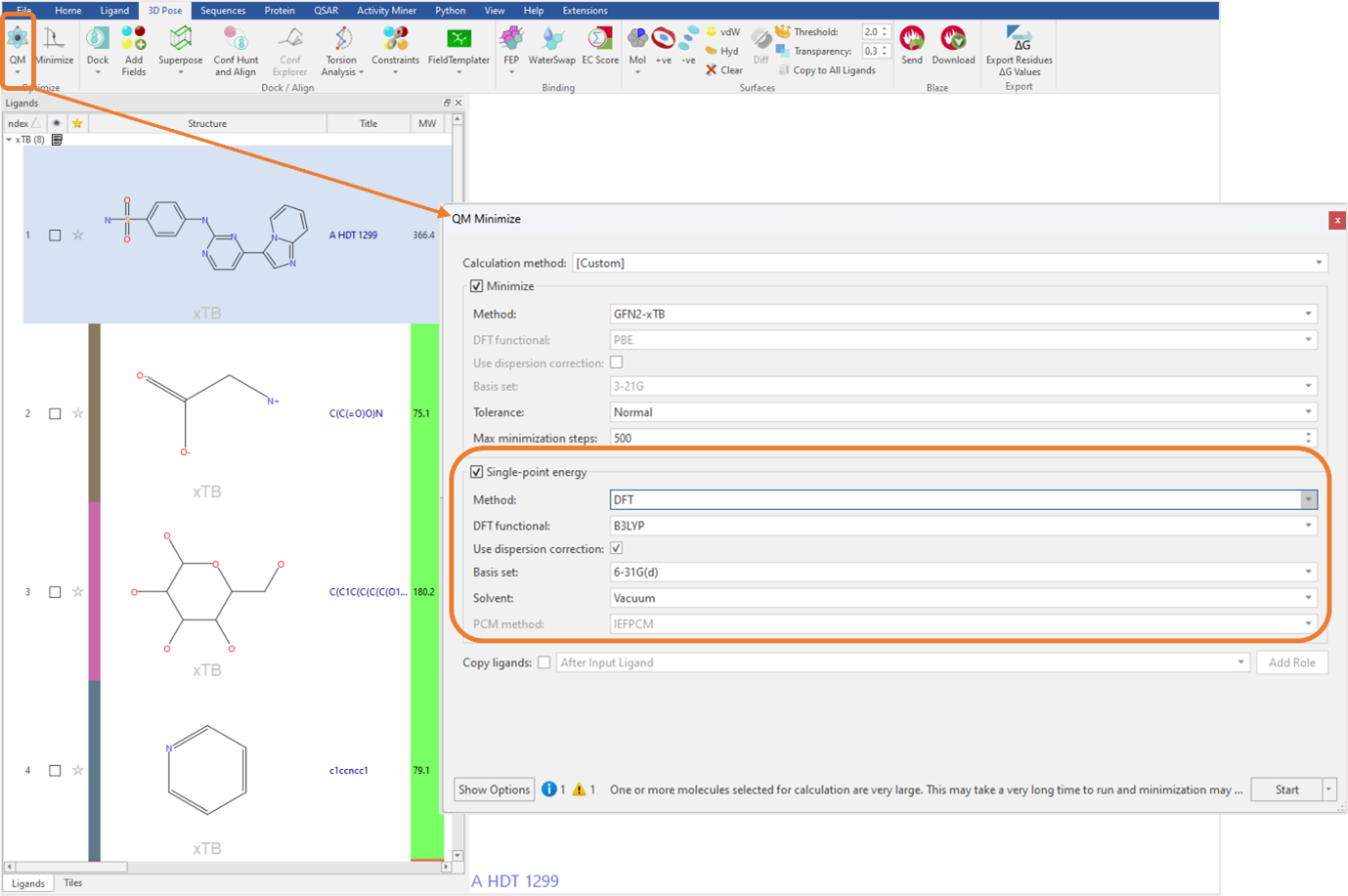
Figure 3: A hybrid GFN2-xTB//DFT workflow, in which the minimization is performed at GFN2-xTB level of theory, followed by a more accurate single-point energy calculation at DFT level of theory.
The GFN2-xTB calculation workflows outlined in Figure 2 and Figure 3 showcase a computationally tractable method for accurately computing minimized structures and energetics for large ligands that fall to the right of Figure 1.
To exemplify this, we can re-create the compute time vs. molecular weight graph shown in Figure 1, but now using the hybrid GFN2-xTB/DFT workflow outlined in Figure 3. As can be clearly seen in Figure 4, the GFN2-xTB minimization and single-point energy computation time for very large molecules is on the order of minutes and not days when plotted relative to the DFT minimization and single-point energy compute time.
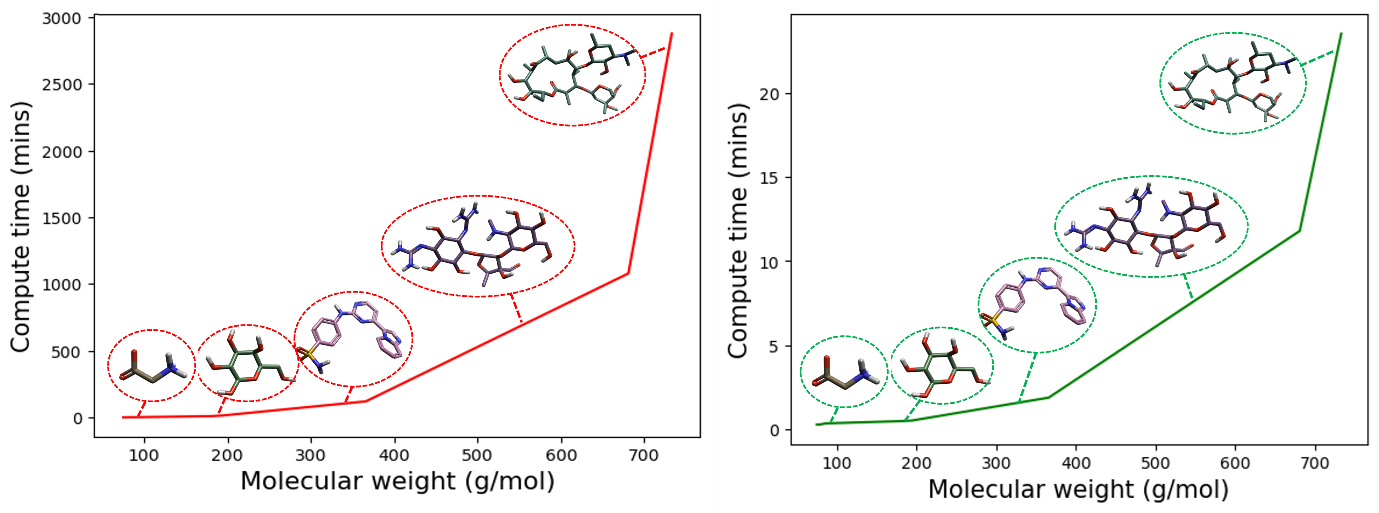
Figure 4: GFN2-xTB//DFT minimization and single-point energy hybrid workflow compute time (mins) as a function of increasing molecular weight (g/mol) plotted in green (right) against the complete DFT minimization and single-point energy calculation time plotted in red, for example biomolecules. From left to right: glycine, glucose, 4-[(4-imidazo[1,2-a]pyridin-3-ylpyrimidin-2-yl)amino]benzenesulfonamide, streptomycin, and erythromycin. The inset displays the GFN2-xTB//DFT compute times plotted in isolation for comparison. The GFN2-xTB//DFT compute times were calculated for an 8-core CPU.
Although GFN2-xTB is a less accurate theoretical approach compared to DFT, the minimized structures predicted by the two methods are frequently in good agreement. For example, minimizing the structure of glucose using both GFN2-xTB and DFT-B3LYP/6-31G(d) (Figure 5), generates optimized structures with a global all-atom root-mean-squared-deviation (RMSD) of only 0.93 Å.
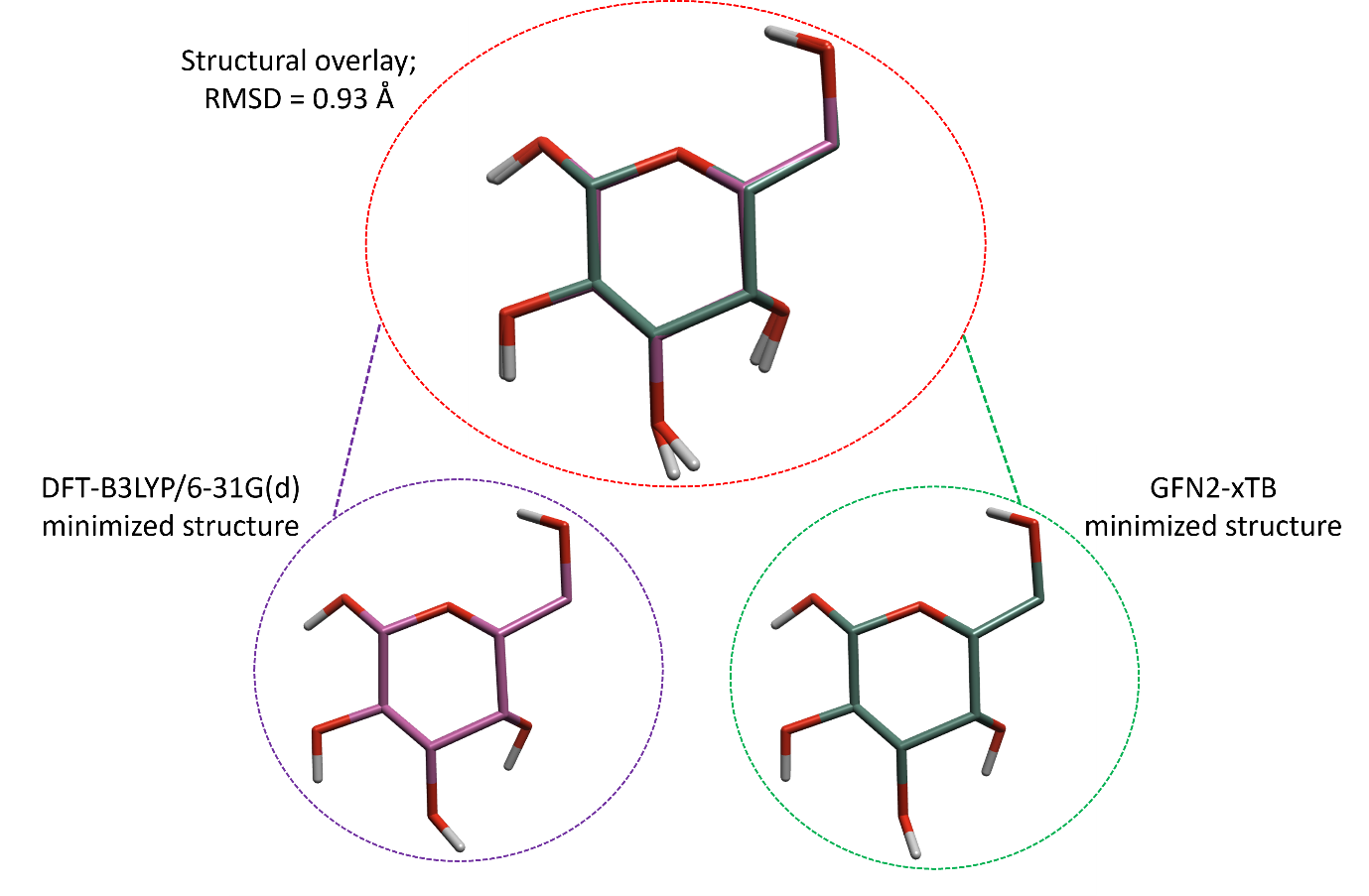
Figure 5: Structural overlay of two glucose molecules, one optimized at B3LYP/6-31g(d) level of theory (purple, left) and the other at GFN2-xTB level of theory (right, green).
In this article, we have highlighted GFN2-xTB as a more computationally tractable method of performing minimizations and single-point energy calculations on larger (100’s atoms) molecular structures. Not only is this method attractive due to the much shorter compute times, but also using a combined GFN2-xTB//DFT hybrid minimization and single-point energy workflow can provide an optimal balance between speed and accuracy.
Request a free evaluation of Flare today to further explore its full portfolio of molecular modeling capabilities.
1. http://www.psicode.org/
2. Bannwarth, C.; Ehlert, S.; Grimme, S. GFN2-XTB—An Accurate and Broadly Parametrized Self-Consistent Tight-Binding Quantum Chemical Method with Multipole Electrostatics and Density-Dependent Dispersion Contributions. J. Chem. Theory Comput. 2019, 15 (3), 1652–1671. https://doi.org/10.1021/acs.jctc.8b01176.
3. https://xtb-docs.readthedocs.io/en/latest/setup.html