Improving PROTAC properties via single-point changes to linkers
We explore how computational methods can be applied to proteolysis targeting chimera (PROTAC) design, to effectively tackle some of the ...
News
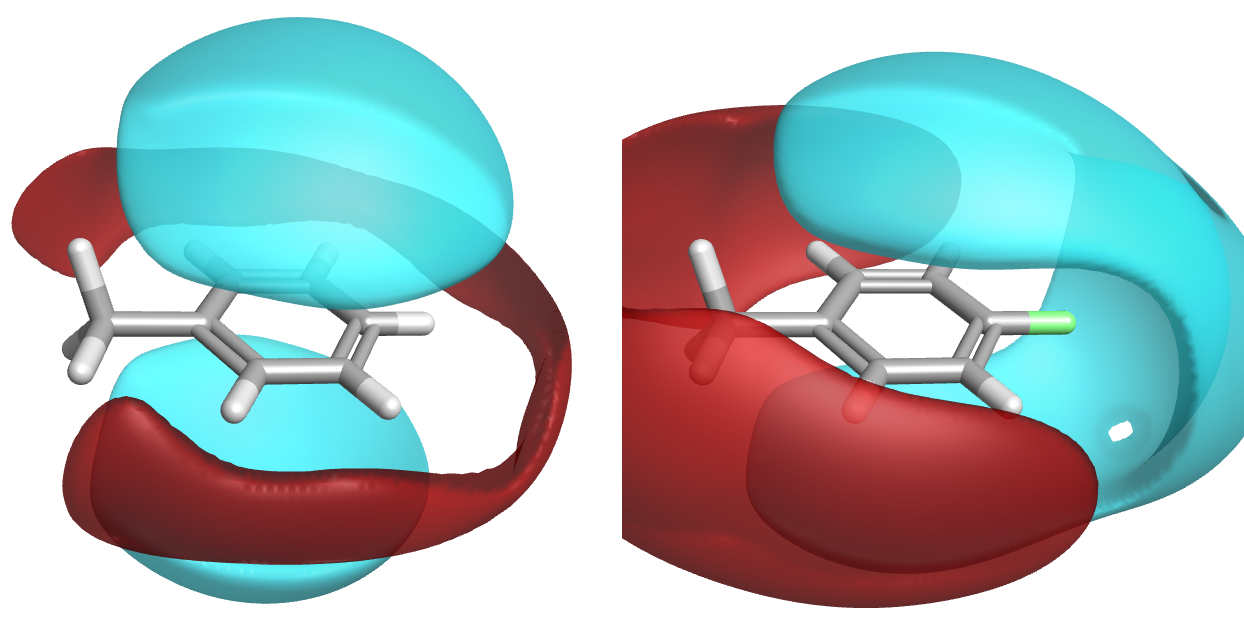
A fundamental problem of SAR interpretation is that what seems a relatively simple change in the structure is rarely a simple change from the protein’s point of view. This is something that was highlighted by The Curious Wavefunction as the fundamental philosophical dilemma of chemistry – it is nigh on impossible to make a single change to a molecule and understand the effects of that change. A simple example I often use is a change from an unsubstituted phenyl to the corresponding 4-fluoro-substituent (Figure 1). The change of hydrogen to fluorine introduces negative charge at the changing atom but also withdraws electrons from the ring, adds a dipole, and increases the positive charge on the aromatic hydrogens. So one atom has been changed, but any change in activity could come from any combination of three distinct interactions.
A real world example can be seen in a series of PERK inhibitors from GSK. The simple change of a pyrrolo-pyrimidine to a furo-pyrimidine causes a loss of 1.3 log units of activity (Figure 2a). Using Activity Miner in Torch or Forge this change shows up as an activity cliff. You can plot the field difference for this pair of molecules – the regions where the molecular interaction potential is greater for one compound than for the other. It shows (Figure 2b) that the relatively modest structural change causes a number of changes to the electrostatics, each of which could contribute to the change in potency.
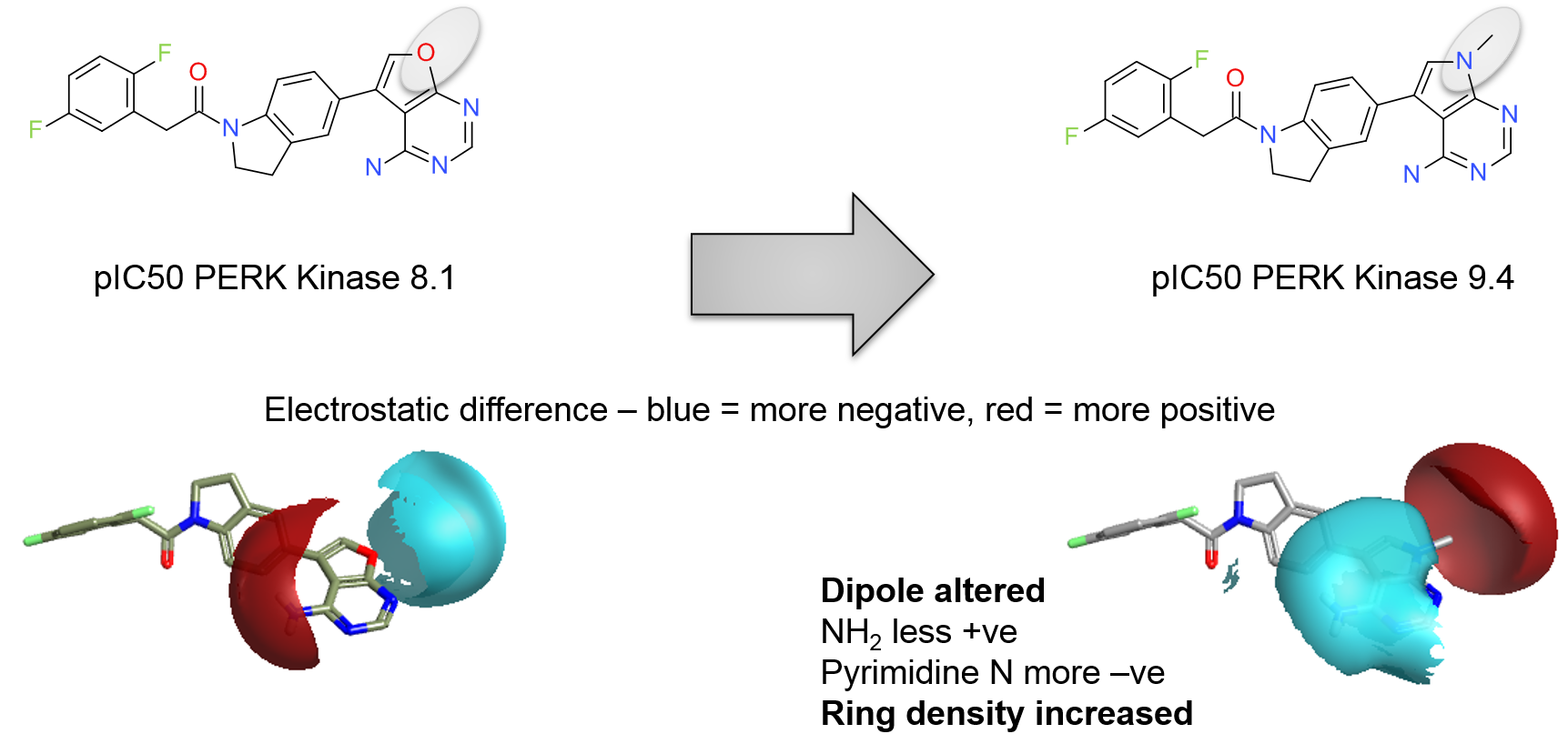
The region that is changed between this pair of ligands (N-Me to O) is solvent exposed and makes no direct interactions with the protein. The large change in activity must therefore be due to the indirect effects. The main ring system is involved in a pi-stacking interaction with Phe943 (numbering from PDB: 4G31) and hence it is quite possible that the changes in the electronegativity of the ring is at the root of the observations. The pyrrolo-pyrimidine is a more electron-rich ring system and hence will make a stronger pi-stacking interaction in this geometry. Examining just the local effects of the structural change in this case thus misses some of the important SAR information.
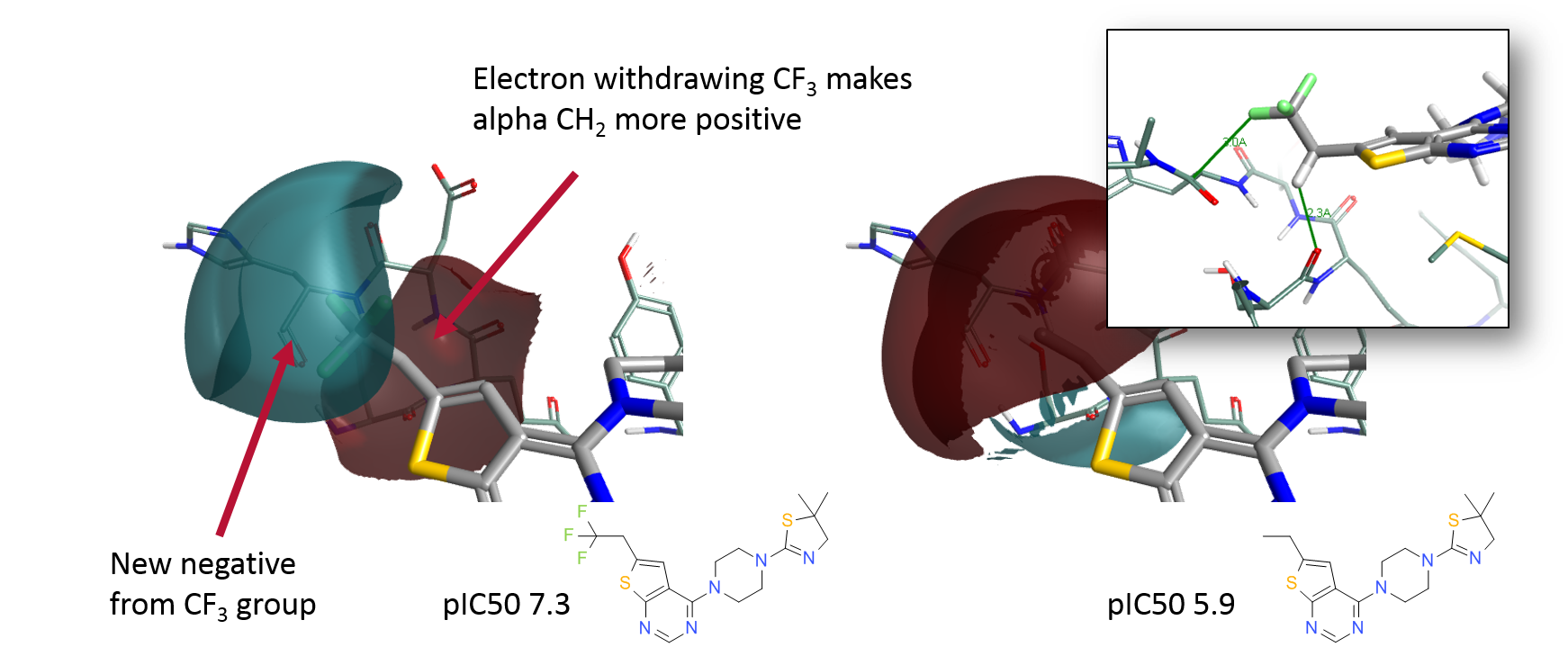
A further example of chemical changes being non-local came to light recently. Pollock et al. were studying the inhibition of Menin and found that replacement of a propyl group with a trifluoroethyl led to a surprisingly large change in activity. X-ray data revealed an interaction between the introduced fluorine atoms and the carbonyl carbon of a backbone amide. They searched the PDB and found that this was a commonly observed interaction and with QM showed it to be favorable, so settled on this as the explanation for the activity change. However changing a methyl to a trifluoromethyl results in more changes than just introducing a negative atom as can be seen if you model the two compounds in Forge (using PDB: 5DD9) and display the field difference in this region (Figure 3). Adding a CF3 group introduces favorable electrostatic interactions with the backbone carbonyl but also polarises the alpha-CH2 to be more positive. It is interesting to note that this alpha CH2 sits very close (2.3A) to the carbonyl of Ser178 and hence the polarisation of this CH2 also contributes to the change in potency. Once again, focusing just on the immediate change without considering the effect of that change on the wider context of the molecule leads to important SAR information being missed.
In many cases the only solution to deciphering the SAR is to make more compounds that steadily test each hypothesis for the underlying causes of activity. However, the release of Activity Atlas models last year has significantly helped in this process. The Activity Atlas method combines changes across multiple molecules and summarizes the regions of important SAR in terms of electrostatics and shape. Activity Atlas does not necessarily require single point changes to develop a meaningful model and hence can add value in complex datasets. Naturally some changes are correlated – in the PERK set above the Activity Atlas model indicates that additional bulk on the methyl of the pyrrolo-pyrimidine is advantageous but this is because all N-Me containing compounds are more active than the O or S analogues and the analysis cannot detangle the correlated steric and electronic effects. However, Activity Atlas definitively shows for this data set than a more electron-rich heterocycle is favored, an important point which might have been missed without this view of the data. More importantly, the model provides a testable hypothesis that can be used to design new compounds. These in turn can be used to update the model, progressing the project knowledge and achieving the project goals in a shorter time.
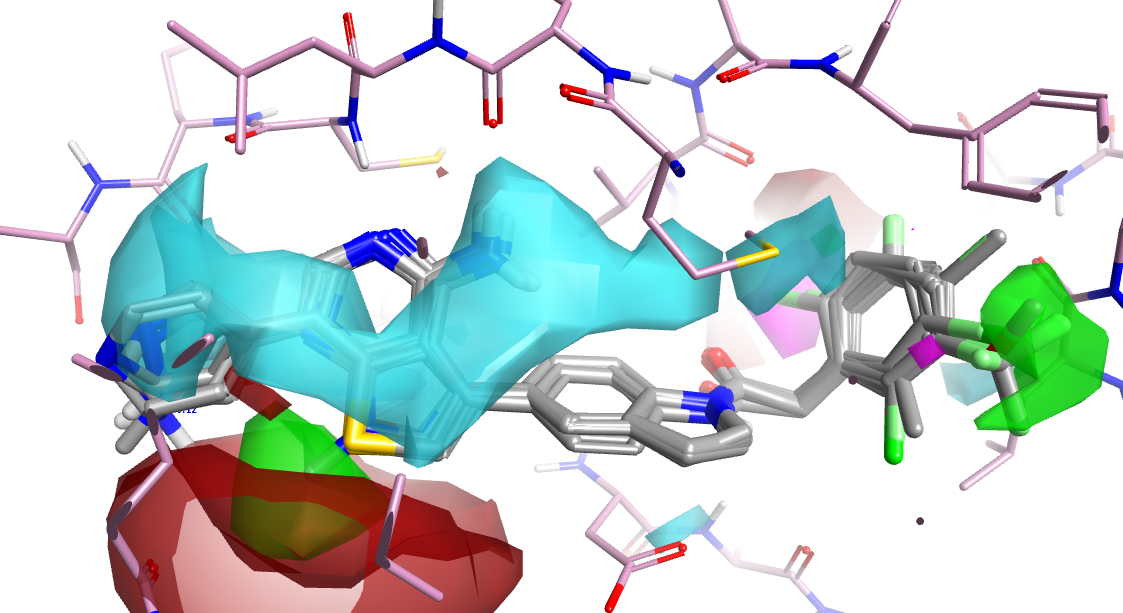
Figure 4. Activity Atlas model from 25 compounds with activity data against PERK. Negative (blue) regions above the pyrrolo-pyrimidine ring suggest activity is enhanced by electron rich heterocycles. Favorable shape regions are highlighted in in green, positive regions in red.