Improving PROTAC properties via single-point changes to linkers
We explore how computational methods can be applied to proteolysis targeting chimera (PROTAC) design, to effectively tackle some of the ...
News

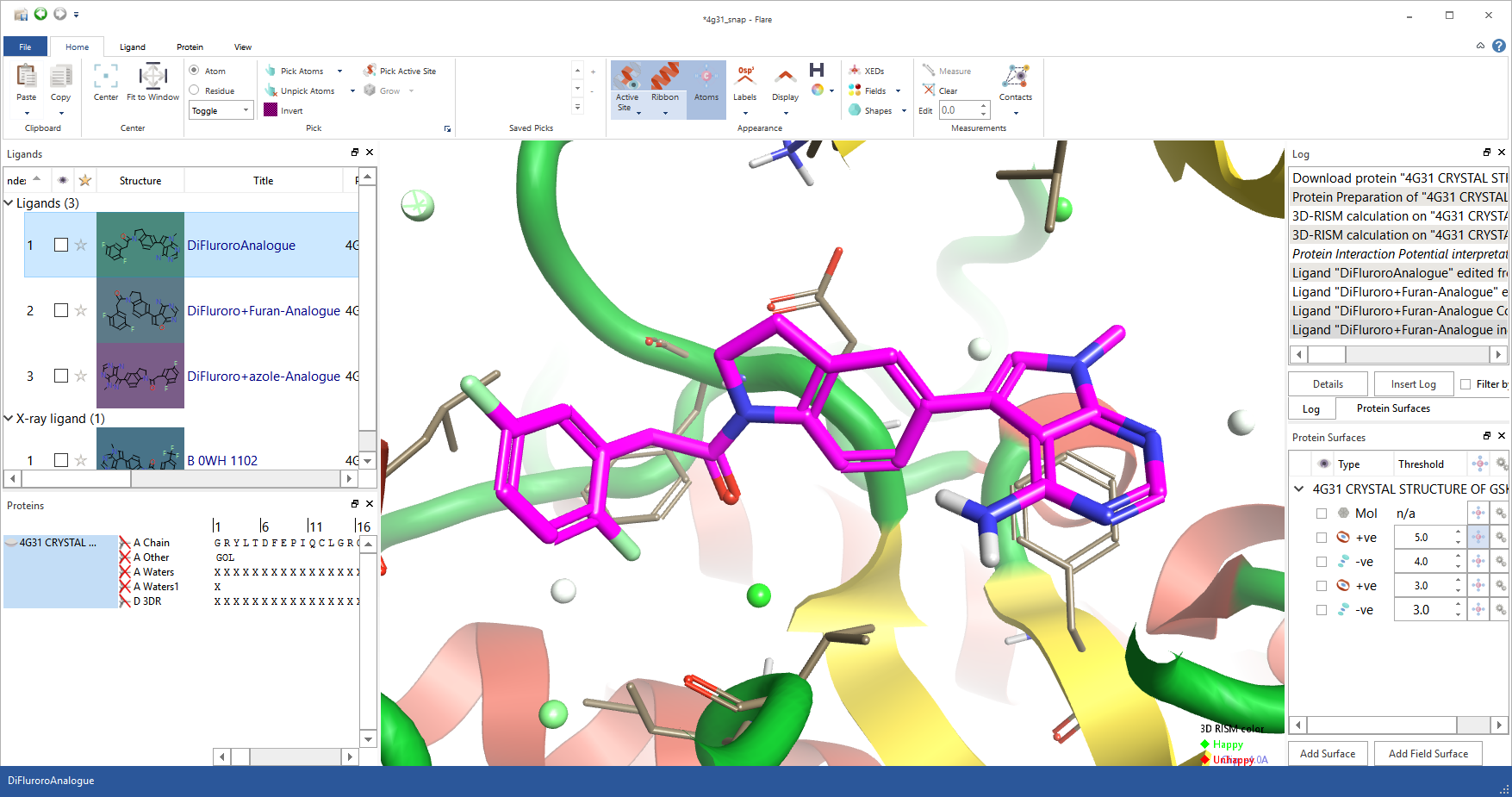
Flare has been created with ligand design at its heart so you can easily navigate ligands and their proteins, comparing, contrasting and improving them. To do this the ‘Molecules’ table has been borrowed from Forge and Torch. The table holds ligands and their data, and has been enhanced with a separate table for proteins. Why two places for molecules? We felt that separating the two types of molecule has distinct advantages. First it enables you to store and display, next to each ligand, all the physico-chemical property data that chemistry designers need to assess designs for progression to synthesis. It enables separate, rapid control of which elements are displayed in the 3D window – for example, you can quickly create a grid and compare one ligand in many different proteins or many different ligands in one protein. Lastly, separating the ligands into their own table enables separation and navigation of ligands in a way that would otherwise not be possible.
To counter any lack of functionality in separating proteins and ligands, drag and drop between the tables has been enabled. To move a ligand into a protein, or separate it away, you simply drag the molecule from one table to the other. Equally, each ligand has a concept of its parent protein and hence it will be associated with the correct protein when viewing multiple ligand protein complexes.
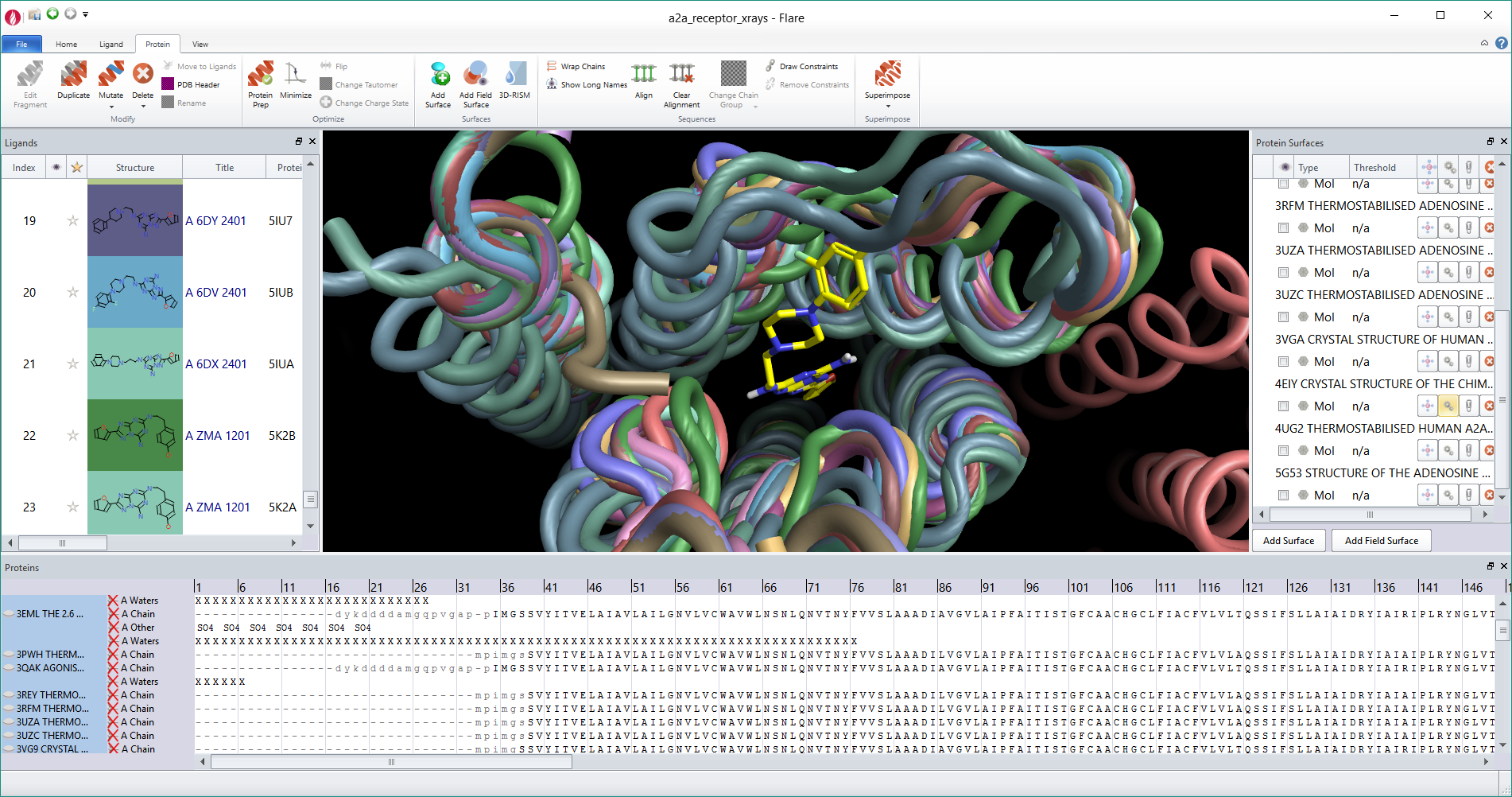
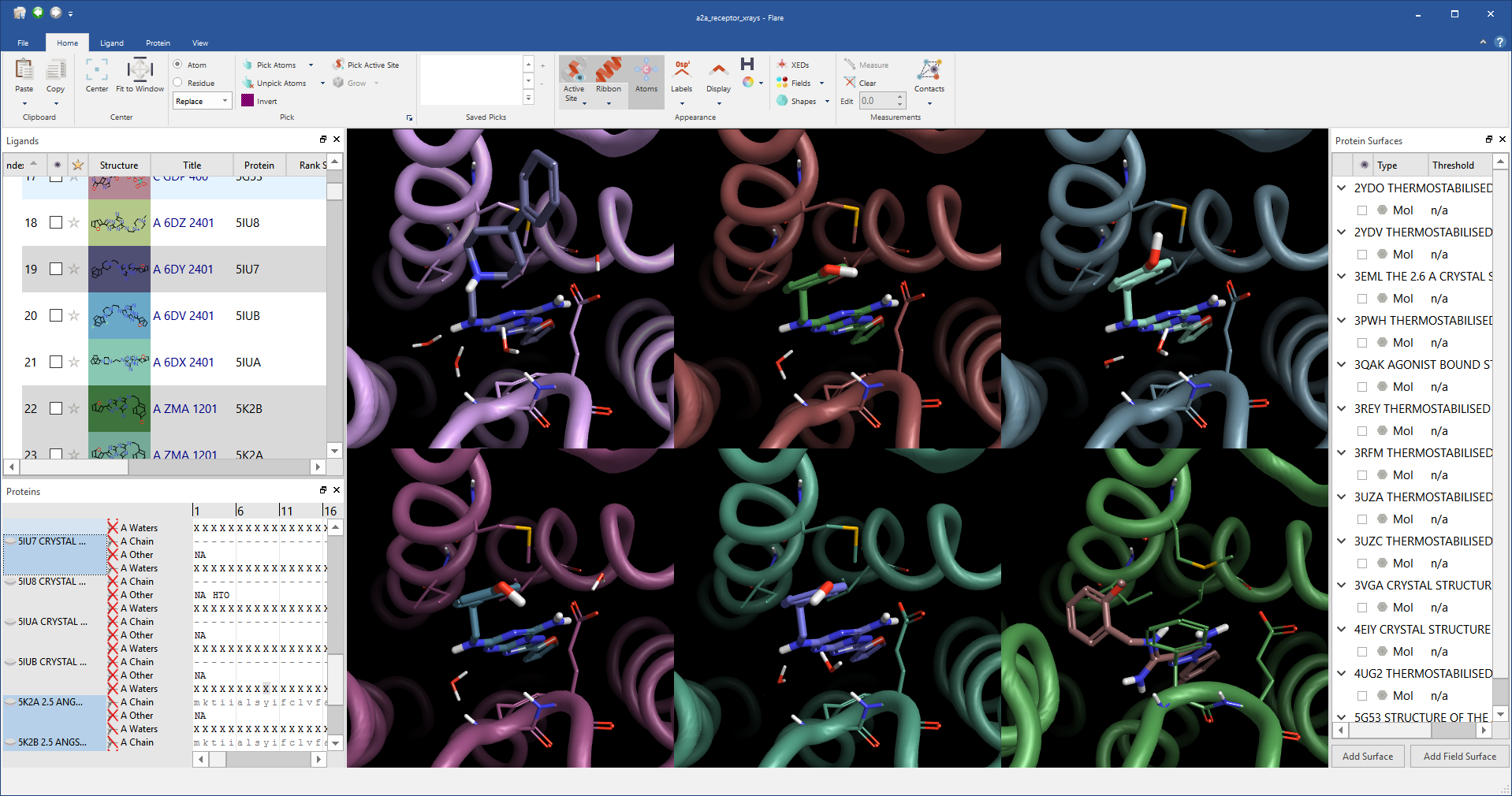
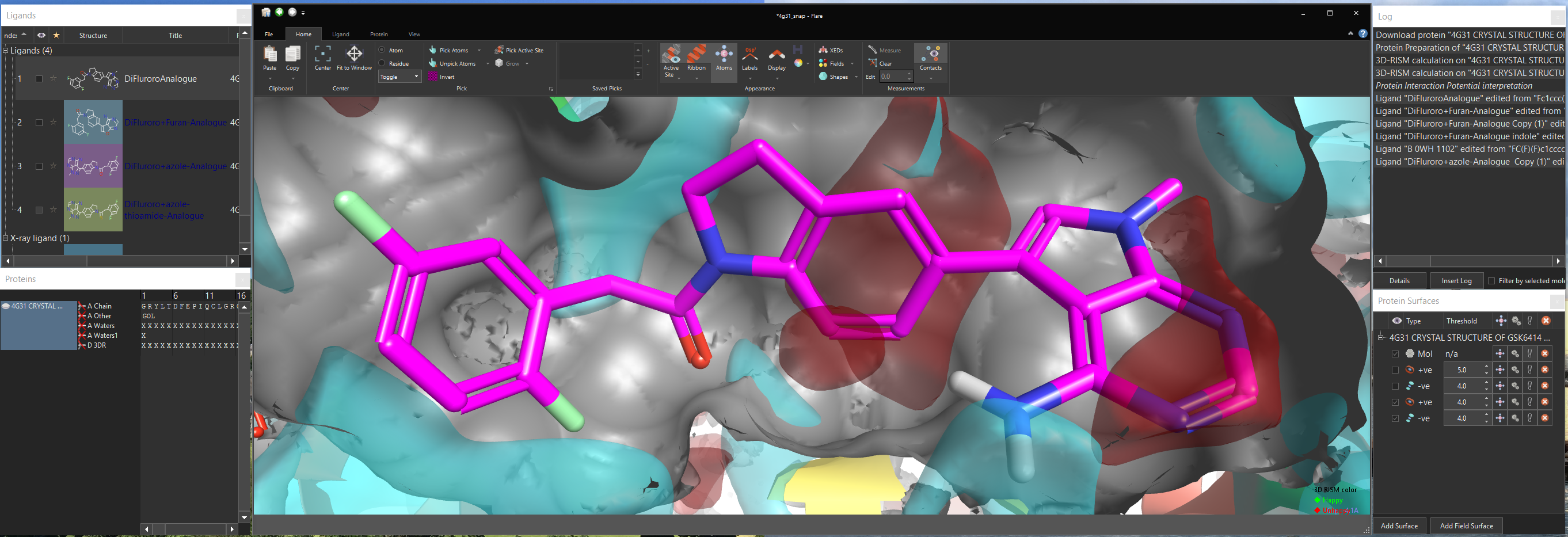
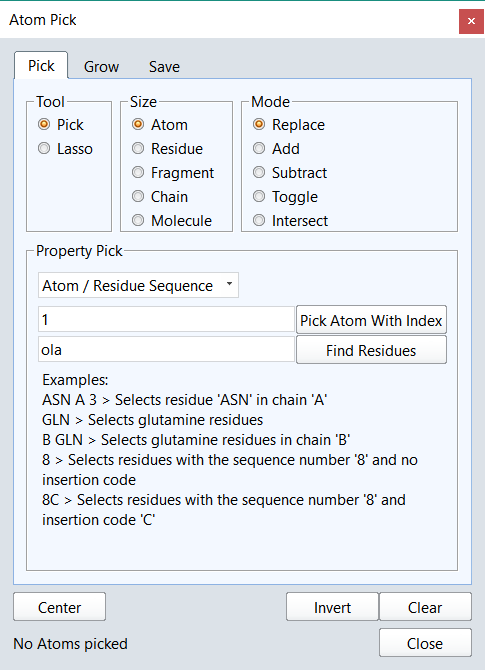
Picking atoms, whether to change the display style, add a surface or perform a minimization is an amazingly frequent action in structure-based design. We wanted to make it as easy as possible, so common picking actions such as picking the active site or all ligand atoms are available directly from the ‘Home’ tab of the ribbon. However, this is just a small selection of the actions in Flare as they are enhanced through an extension, accessible from the ribbon, which gives a depth of functionality to Flare’s picking algorithms. For example using the extension you can pick atoms based on a SMARTS pattern, pick residues using a text query such as ‘ASN 83’, chains by name, residues by names or numbers, add or subtract to the existing pick or take the intersection. Using the enhanced picking widget you should be able to grab any atom within the application without needing to first find it in the 3D window.
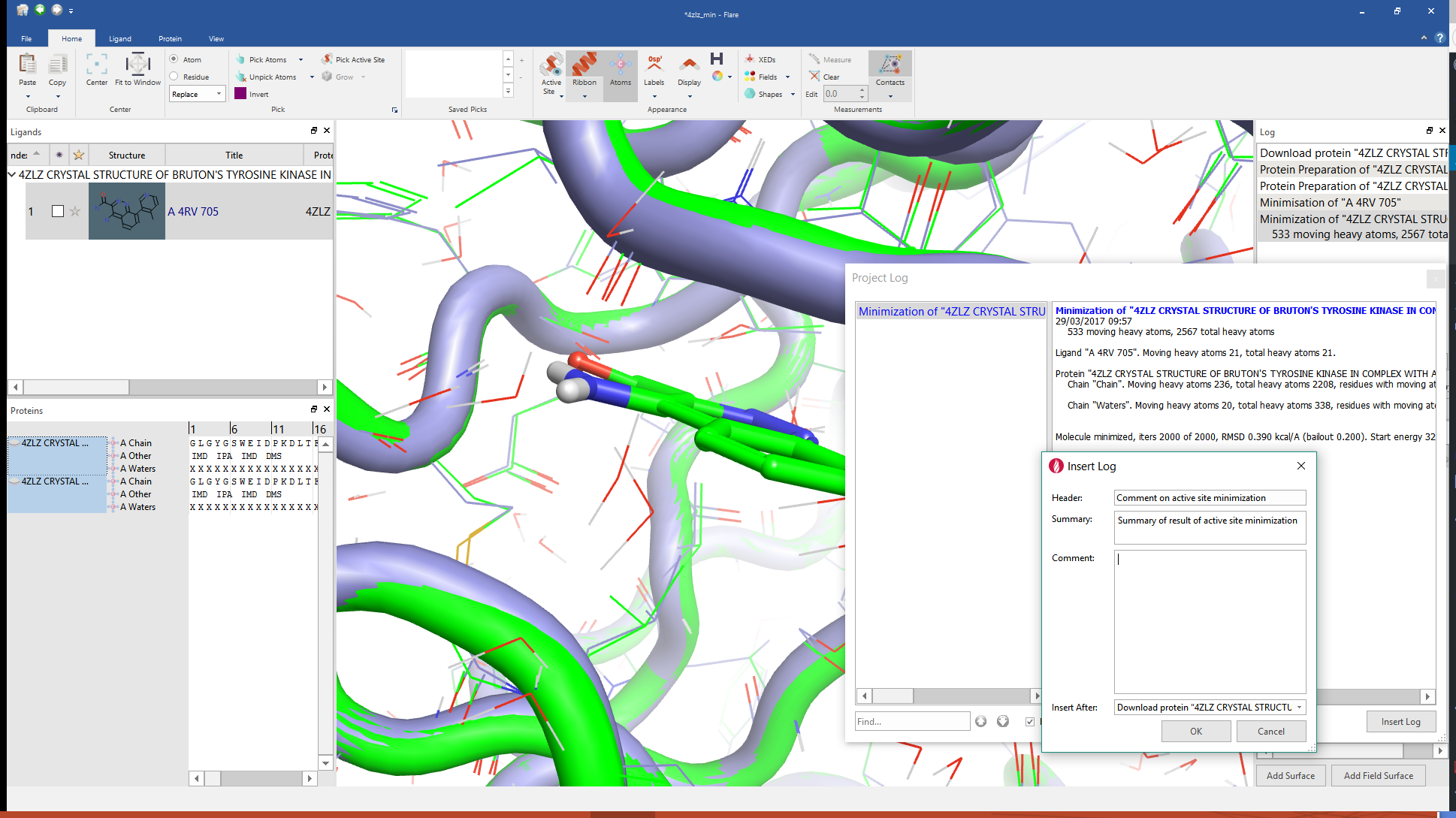
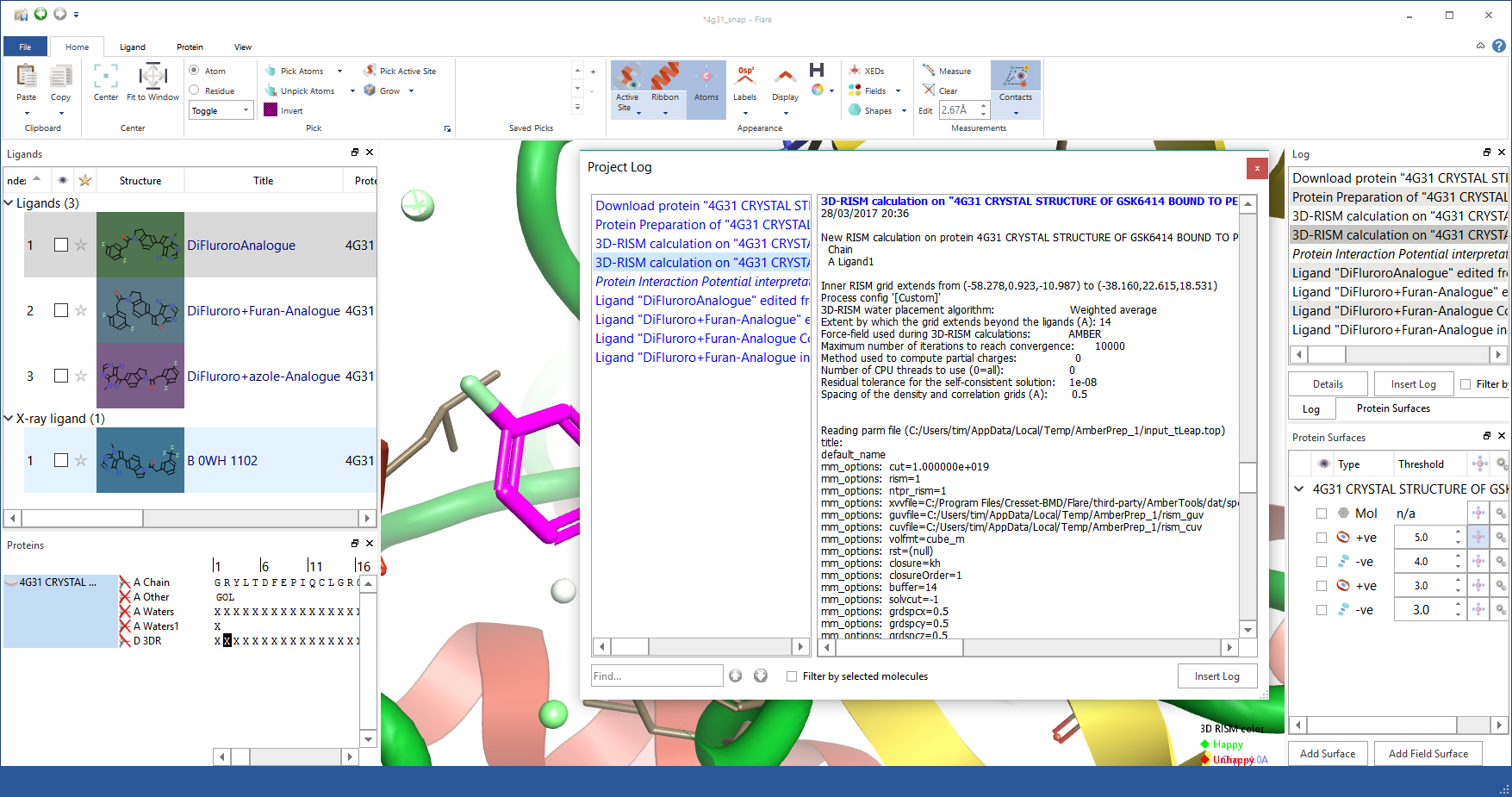
A key piece of feedback from alpha and beta test phases was that you wanted detailed logging. To get the right balance between finding the relevant information and seeing the detail of the step there is a hierarchy of logging. All top level events are recorded to a log window that you can choose to keep visible, move to the side or close as you prefer. At any time if you want the detail behind an operation then you can go to the log window and double click the relevant entry to see all the detail that underlies the operation in question.
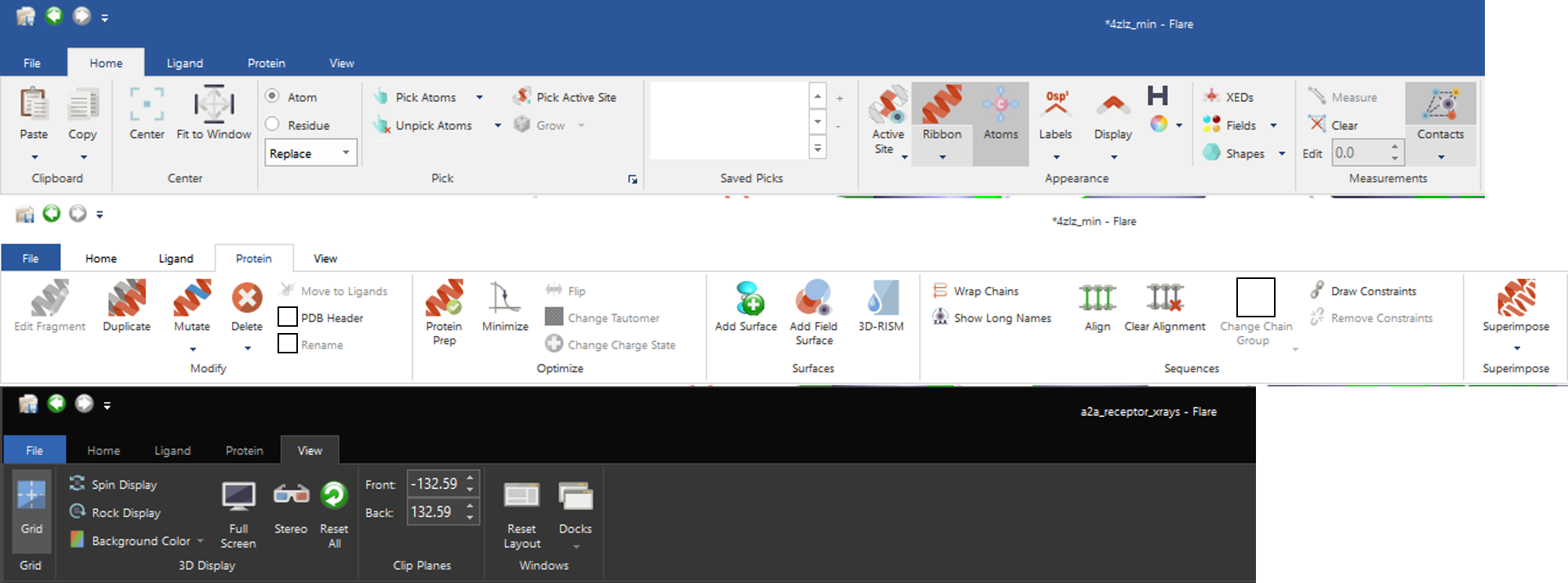
Our intention is for Flare’s capabilities to grow significantly over time so we have built a GUI with room to expand the command structure without compromising usability. A key element is the choice of a ribbon interface instead of traditional menus; these provide a logical framework for commands with an easy search strategy to find the one that you need at that moment. We were always mindful to enable customization in the fullness of time and enable users to control their own work environment and the ribbon interface is the perfect environment for this. Our intention here is to avoid the nightmare growth of multiple, unexplained and unobvious icons suffered by many applications and classically described in the story of the microsoft ribbon.
Flare will be available for evaluation very soon. If you would like to test drive the novel interface, or apply one of the novel scientific methods to your project, please contact us to register your interest.