The adaptability of pyFlare to access advanced data visualization and calculation functionalities
We showcase some examples within a drug discovery context where the pyFlare environment has been further expanded with advanced data ...
News
The next version of Spark (V10.3), Cresset’s bioisostere finder and idea generator, will be released in the next couple of weeks. In this sneak peek we take a look at a few of the exciting new features that have been requested.
In V10.2 we introduced radial plots that give a quick summary of the physical properties of compounds but they can sometimes be lost in the detailed results and clustered results views. Since then customers have told us that they really wanted a simpler view which focussed on the structure of the bioisostere with the option to include the radial plot or selected data. In V10.3 we satisfy all these requests with the new tile view of results (below).
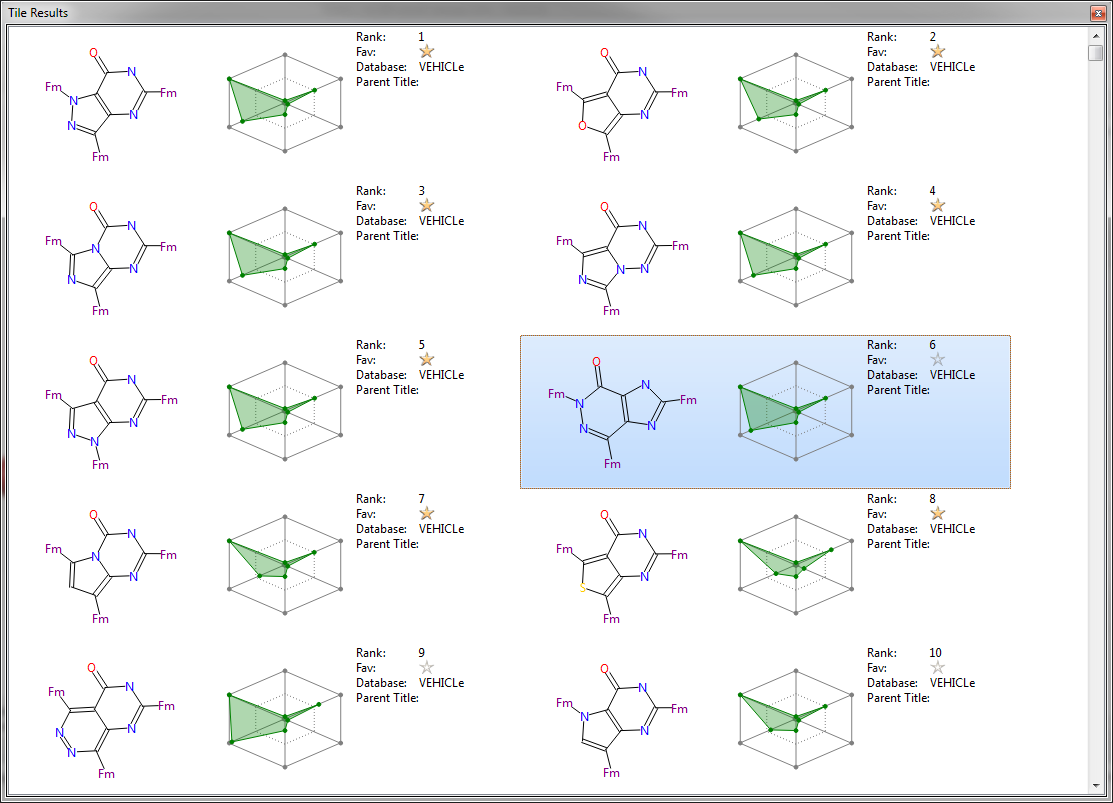
The tiles come in three sizes – large (shown above), medium and small. They can have any amount of data displayed on them. For example, the view above shows the rank, favorite status and originating database for a fragment together with the radial plot for each result which summarizes the physical properties of each compound compared to ideality (perfect results appear with a small area near the center). This can be customized to just show exactly the information that is most useful to you.
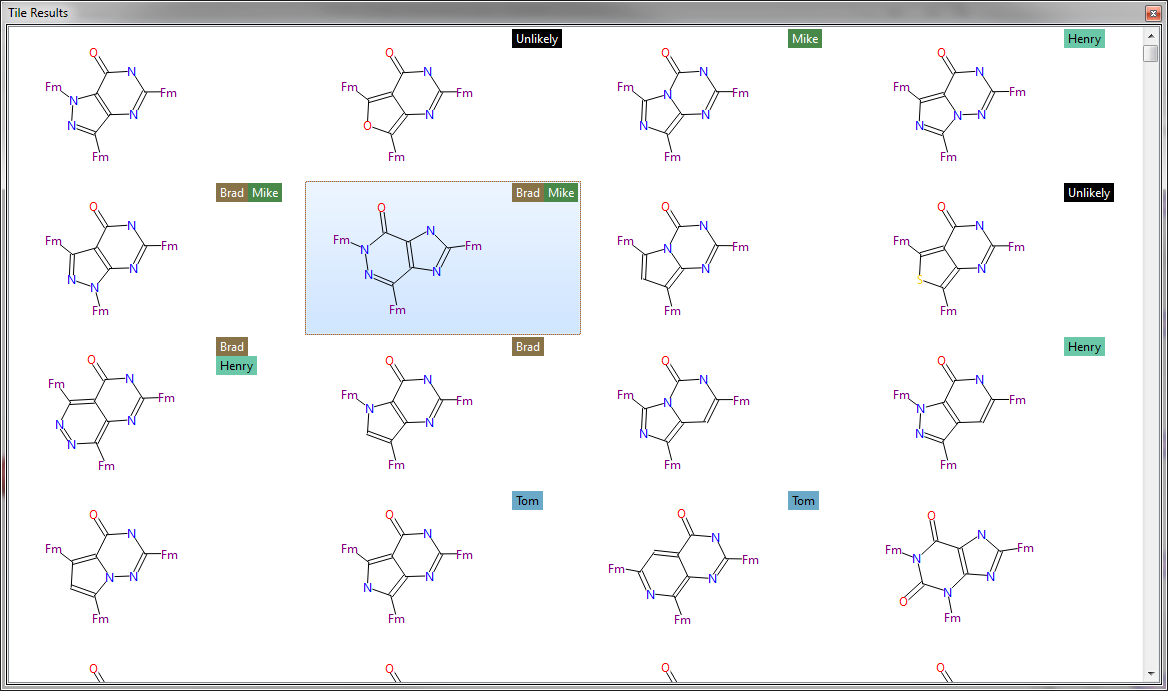
In this release we bring in the idea of result tags so results can be labelled in a more finely than using the favorites label. A tag is an arbitrary piece of text that is attached to a result so it can be used in any way that helps you find the bioisostere that will ignite your project. For example, tags can be used when sharing the results of an experiment with colleagues where each person tags the results that they like with their name. In this case you can simply apply filters on the tags to see the results that everyone likes or to investigate each individuals choice.
The features above are just the tip of the iceberg for Spark V10.3. We have many more features to show in you in the full release announcement. These include SMARTS filters on database searching and on results tables, custom notes on results, improved importing from pdb files, improved protein display, improved display of protein interactions, a new wizard, updated databases, improved database downloader, new databases, increased information about the source of a fragment.
To take advantage of all these new features we encourage you to update as soon after release as possible.
Contact us to arrange a free demo of the new features.