The adaptability of pyFlare to access advanced data visualization and calculation functionalities
We showcase some examples within a drug discovery context where the pyFlare environment has been further expanded with advanced data ...
News
A new version of Spark, our scaffold hopping and bioisostere replacement tool, is now released. V10.4 includes many new or improved features and gives access to new and updated chemical diversity.
The development of our applications is guided by our customers and this release is bursting with new features and science that you have asked for. A few of these are described below, however, I suggest that you use the software for yourself to discover the other features and see them in action.
The new Spark reagent databases are derived from eMolecules’ building blocks and replace the previous reagents based on ZINC. These new databases enable Spark users to select the most promising results from their experiment with confidence that the corresponding reagents will be commercially available from reliable suppliers and access up-to-date availability information.
The chemically intuitive rules for R-group database creation have been refined to improve the accuracy of the chemistry incorporated into the new reagent databases. Over 20 different reagent databases are provided by Cresset using the updated rules, which can be easily modified to suit your preferences. If you think something is missing then let us know and we can add it to the list in minutes.
Customers with a database generator license can use our rules to process their own available reagents, giving rapid suggestions for the next set of compounds to be made using the reagents currently in your lab.
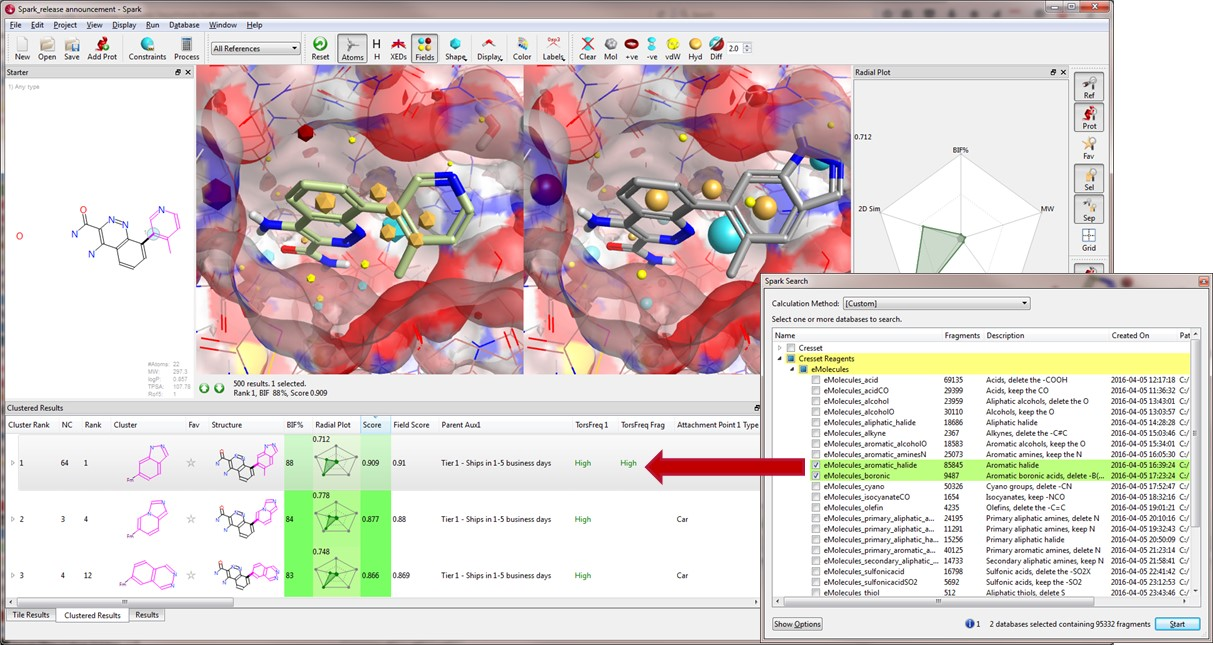
New Results table columns are available within Spark V10.4, to facilitate the analysis of the quality of the results obtained from your Spark experiment and the assessment of chemical feasibility.
The ‘TorsFreq Frag’ and ‘TorsFreq’ column values are computed by analysing the frequency of torsions, as recorded in the CSD, using the Torsion Library method jointly developed by the University of Hamburg (Center for Bioinformatics) and F. Hoffman-La-Roche Ltd. The analysis is carried out for all dihedrals associated with rotatable bonds within the bioisosteric replacement and for each new bond formed in the result molecule. Torsions associated with a low frequency are a possible cause for concerns and should be further investigated.
Spark has always enabled you to restrict your search to fragments that link through a specific atom type. This feature enables you to search for bioisosteres that would work with your synthetic scheme. In this release we have added the ‘Attachment Point Type’ into the main result sheet to enable you to perform a wider search and then focus on the results that are of interest to you. This facilitates the assessment of chemical feasibility, enabling you to focus on those results which match the chemical strategy you have in mind for your project.
One of the most requested features by customers is the ability to include corporate or externally-computed data for any compound into the Results tables. Spark V10.4 can connect to an external web service, through a REST interface, to import external properties and data computed or retrieved by such web services as additional columns in the Results tables. Using the new service you can bring in external predictions for new designs or simply use the corporate algorithm for calculating logP. Once imported the properties can be used in the Radial Plot, Tiles View and for coloring molecules and table cells enabling you to monitor the overall property profile of the results your Spark experiment.
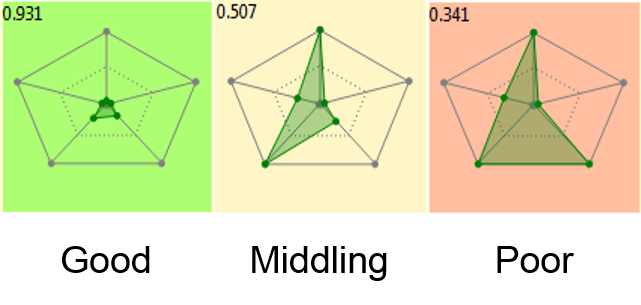
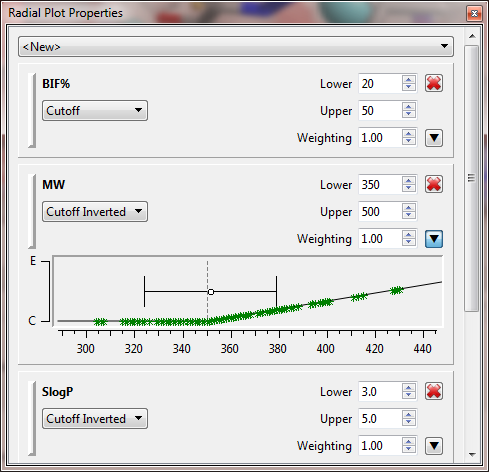
Radial plots were introduced in SparkV10.3 to provide a graphical representation of numerical data. These initial radial plots created a simple picture to show how a molecule fits the physicochemical profile of a project with the idea that parameters are within an ideal range, an unacceptable range or somewhere in between. In this release this representation is enhanced by introducing the option to combine all the scores in the radial plot together into a single number scaled between zero and 1 that represents how well the result molecule fits your project profile (Figure 2).
This release represents a significant improvement in the usability and flexibility of the leading bioisostere application. We encourage you to upgrade your version of Spark at your earliest convenience.
If you are not currently a Spark customer, please download a free evaluation.
Contact us if you have queries relating to this release.