Improving PROTAC properties via single-point changes to linkers
We explore how computational methods can be applied to proteolysis targeting chimera (PROTAC) design, to effectively tackle some of the ...
News
Spark™ V10.6, the new version of our scaffold hopping and bioisostere replacement tool, enables medicinal and computational chemists to generate innovative ideas for their drug discovery projects with new and improved search methods.
The new ‘docking’ feature enables Spark to find fragments picking ligand-protein interactions directly from the protein active site. Use this method to grow ligands and fragments into unoccupied pockets of the target protein, and whenever you wish to find novel results making interactions with the active site of your protein not mapped by an existing starter or reference molecule.
Figure 1 shows the details of this new workflow. Spark starts by removing the portion of the starter molecules which you wish to replace (Figure 1, A-B). A new fragment is then attached to the truncated starter molecule and placed in the active site of the protein in a sensible orientation guided by the protein electrostatics calculated with the Cresset XED force field (Figure 1, C). The pose of the new result molecule is then optimized by using the Lead Finder™ ‘score only’ docking mode (Figure 1, D). Finally, the best results are prioritized by means of the Lead Finder ‘Rank’ scoring function, optimized to correctly reproduce the crystallographic pose of known ligands.
Docking constraints mapping relevant ligand-protein interactions can be easily set to ensure that these are maintained in the result molecules.
Docking in Spark is available as an add-on to traditional Spark bioisostere replacement methods. Customers can contact us to get this add-on.
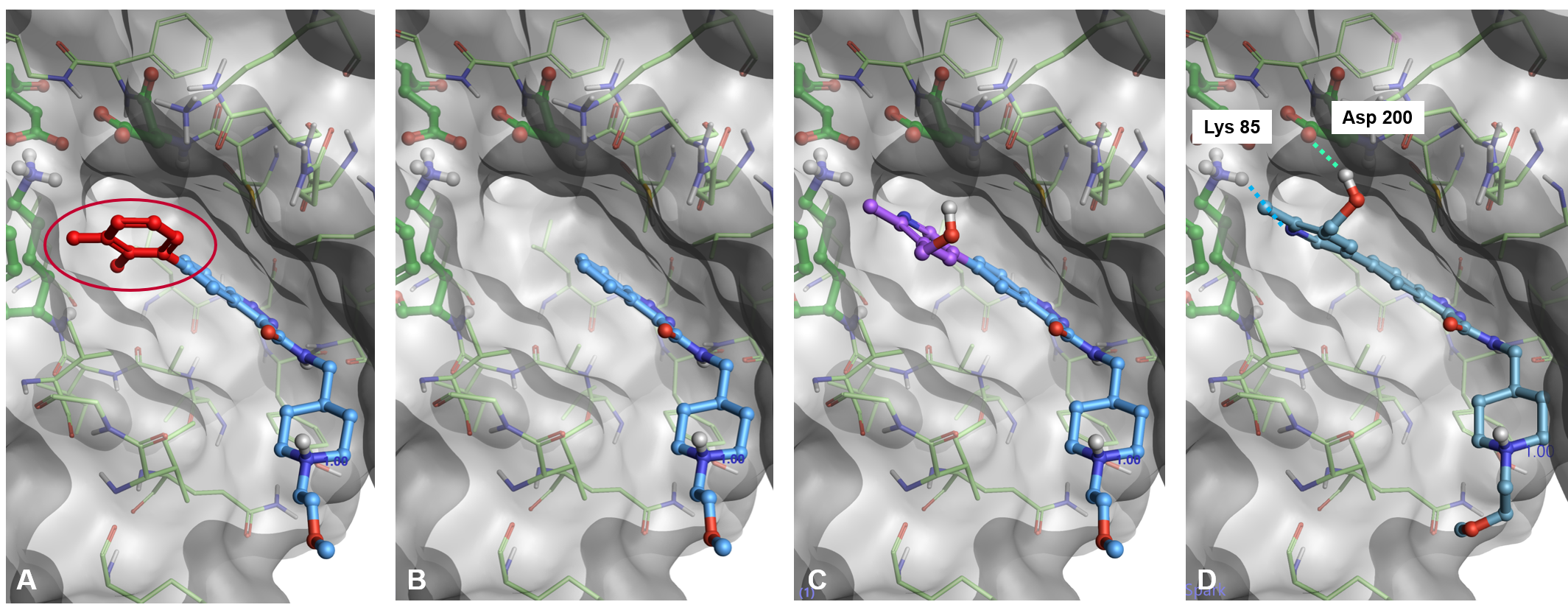
Figure 1. Docking workflow in Spark (PDB: 6TCU). A) A portion of the starter molecule is selected for replacement. B): The selected fragment is deleted. C): A new fragment is attached in a sensible orientation to the truncated starter molecule. D): The pose of the new result molecule is optimized using Lead Finder.
Improvements to the ‘traditional’ Spark methods, based on ligand field/shape similarity, include:
These improvements increase the accuracy of the Spark search on finding interesting bioisosteres. For example, in a scaffold hopping experiment from the X-ray structure of a mineralocorticoid receptor modulator (PDB 5MWP), Spark finds structures which closely resemble other known mineralocorticoid ligands (Figure 2). The results list also shows a modest enrichment in alternative replacements with high Bioisostere Factor. However, in other cases, these improvements can lead to the identification of up to 30% more results with promising BIF% in typical R-group replacement experiments.
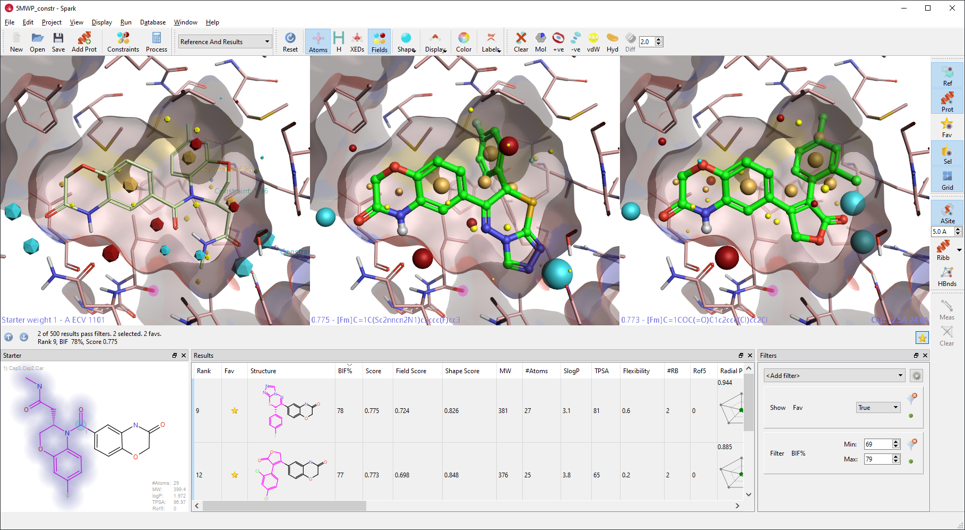
Figure 2: Scaffold hopping experiment on 5MWP finds known scaffolds for the mineralocorticoid receptor and a variety of alternative scaffolds with promising BIF%.
Applying appropriate filters during the search is a great way to reduce the search space for your Spark run, reduce the calculation time, and focus the experiment on the results you really want. This is particularly useful in fragment growing experiments, or when using the docking method, where you want your molecule to grow – but within limits.
To help you with this, the Size Criteria panel of the advanced Spark search options (Figure 3) has been improved. This now reports the number of heavy atoms, MW and number of rotatable bonds for both fragments and result molecules. Use this panel in combination with the information in the Starter window and the Advanced Filters to set the ideal physico-chemical profile for the result molecules from your Spark experiment.
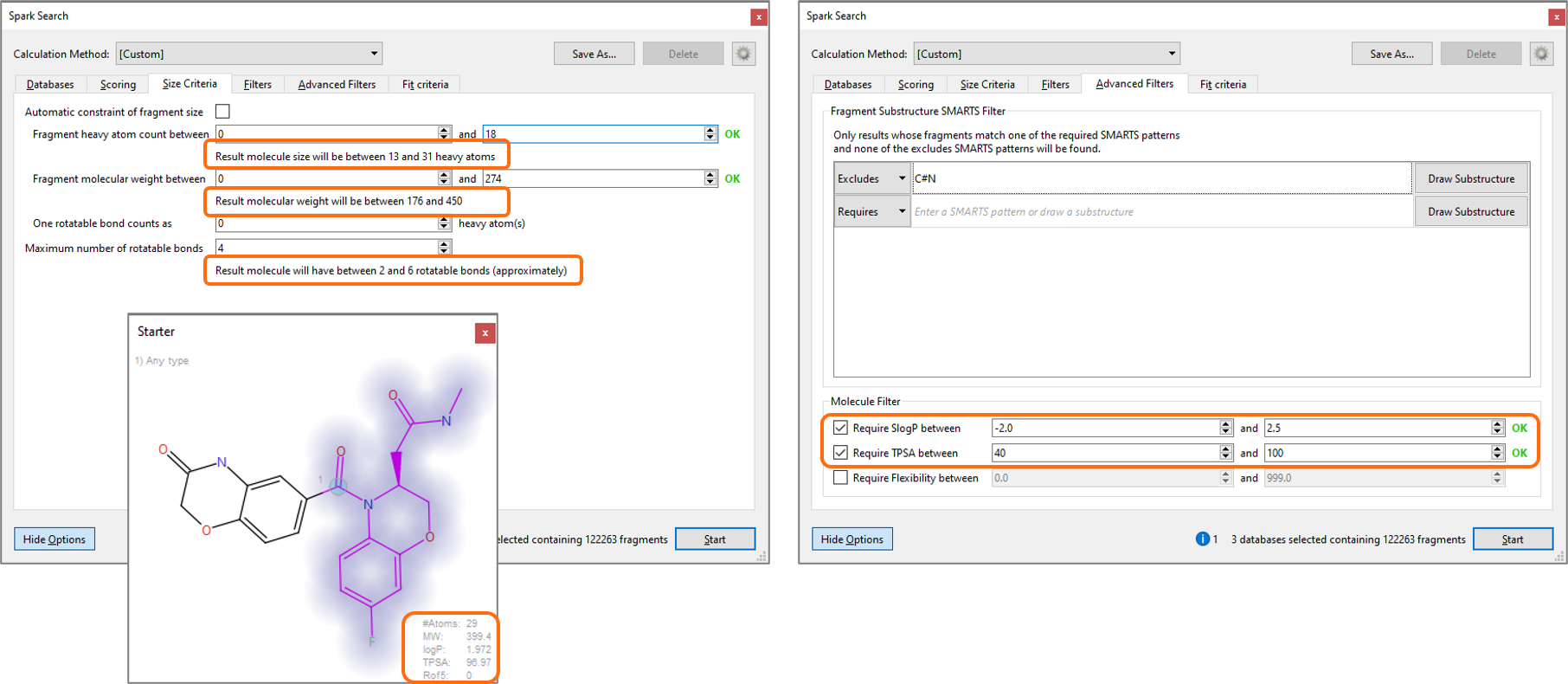
Figure 3: Improved Size Criteria panel (left), used together with the information on the Starter window (bottom) and the Advanced Filters (right), makes it easier for you to set an appropriate physico-chemical profile for the result molecules from your Spark experiment.
This Spark V10.6 release also delivers new science together with significant improvements to the user interface, including:
Current Spark customers will already be in receipt of an email from us with details on how to update to Spark V10.6.
If you’re currently not using Spark, but would like to try it on your project, request an evaluation.