Improving PROTAC properties via single-point changes to linkers
We explore how computational methods can be applied to proteolysis targeting chimera (PROTAC) design, to effectively tackle some of the ...
News
We recently announced our collaboration with BioMolTech, a small modeling software company best known for their docking software, Lead Finder. Cresset has been traditionally focused on ligand-based design, but as we expand our capabilities into more structure-based methods we realized that we would have to supply a robust and accurate docking method to our customers. So, why did we choose Lead Finder?
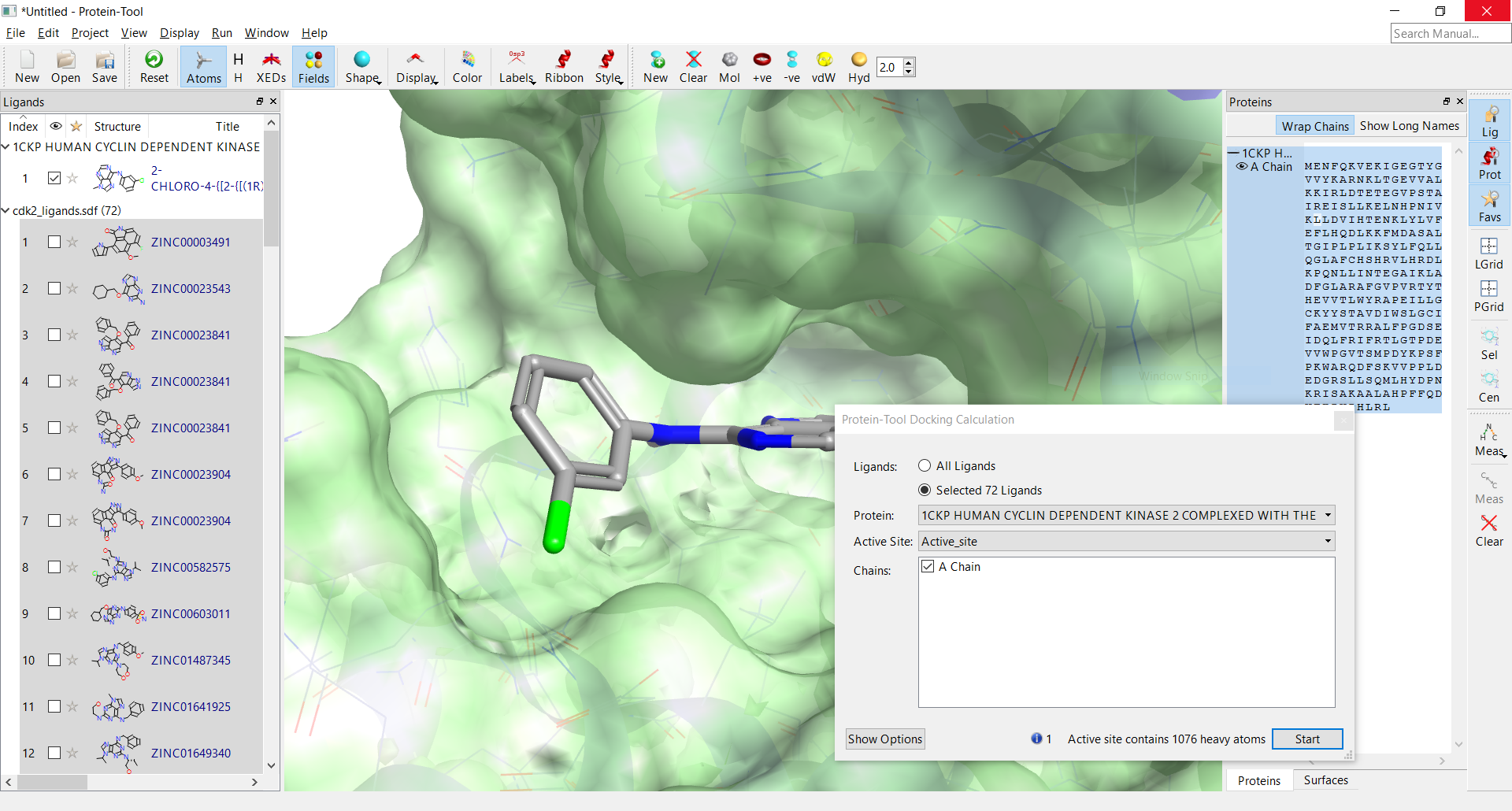
A graphical interface to Lead Finder will be included on our new structure-based design application.
The requirements for a great docking engine are simple to state: it needs to be fast and it needs to be accurate. The latter is by far the most important: nobody cares how quickly you got the answer if it is wrong! Our first question when evaluating docking methods was therefore to ask how good it was. This is actually a difficult question to ask, as there are several different definitions of ‘good’ depending on what you want: virtual screening enrichment? Good pose prediction? Accurate ranking of active molecules?
The first of these, virtual screening, is what most people think of when they think of docking success. Lead Finder has been validated on a wide variety of target classes and shows excellent enrichment rates (median ROC value across 34 protein targets was 0.94), even on targets traditionally seen as very hard such as PPAR-γ. The performance on kinases was uniformly excellent, with ROC values ranging from 0.86 for fibroblast growth factor receptor kinase (FGFR) to 0.96 for tyrosine kinase c-Src.
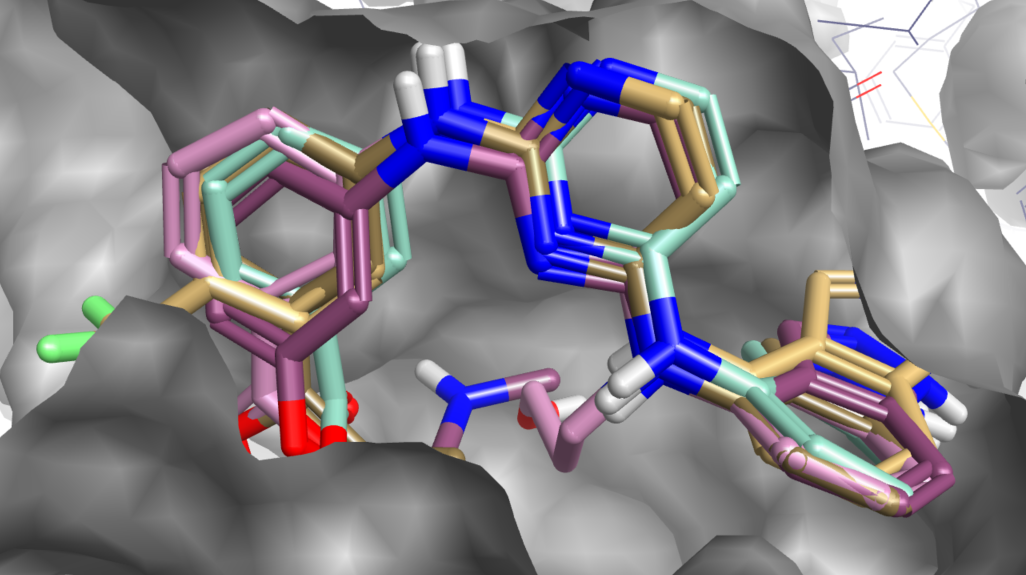
A series of SYK ligands docked to PDB 4yjq with crystal ligand shown in purple.
Pose prediction is of more interest to those working in the lead optimization phase, where assessing the likely bound conformation of a newly-proposed structure can be very helpful in guiding design. Here, too, Lead Finder performs well. On the widely-used Astex Diverse Set, used to test docking performance, Lead Finder produces the correct pose as the top-scoring result 82% of the time, which is comparable to other state-of-the-art methods (Gold, for example, gets 81% on the same measure). On a number of literature data sets testing self-docking performance Lead Finder finds the correct pose between 81 and 96% of the time, which is excellent.
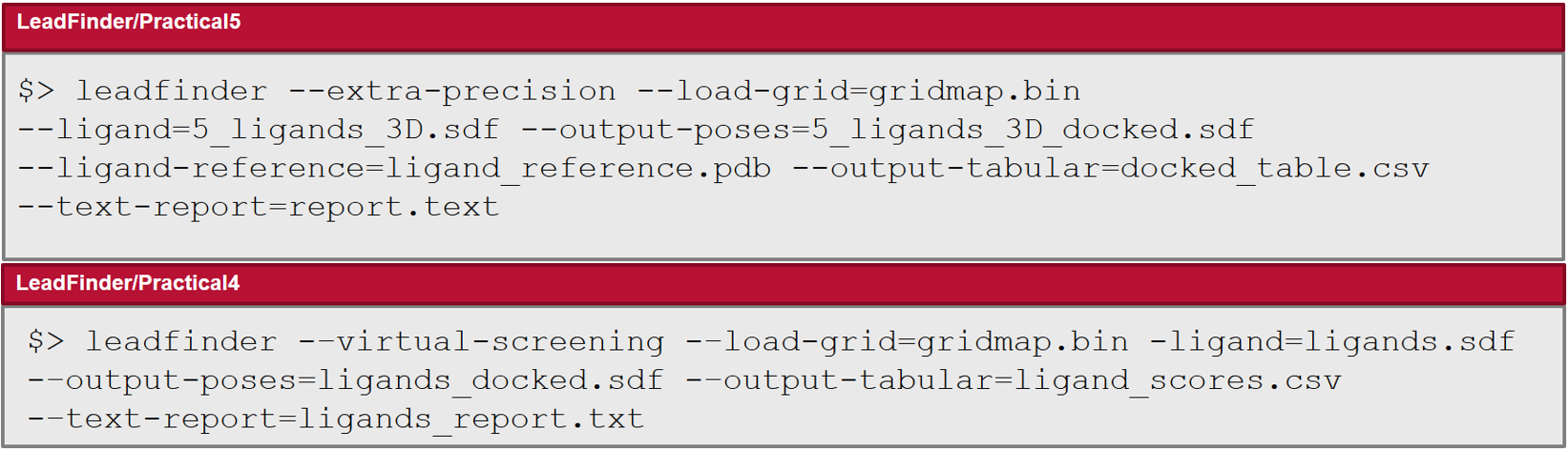

Lead Finder includes dedicated modes for extra-precision and virtual screening experiments.
One of the most intriguing things about Lead Finder is the makeup of the scoring functions. In contrast to many other scoring functions which use heuristic or knowledge-based potentials, the Lead Finder scoring functions comprise a set of physics-based potentials describing electrostatics, hydrogen bonding, desolvation energy, entropic losses on binding and so on. Different scoring functions can be obtained by weighting these contributions differently: BioMolTech have found that the optimal weights for pose prediction differ slightly from those for energy prediction, for example. A separate scoring function has been developed which aims to compute a ΔG of binding given a correct pose. This is a difficult task, and the success of the Lead Finder function was demonstrated in the in the 2010 CSAR blind challenge, where the binding energy of 343 protein-ligand complexes had to be predicted ab initio. Lead Finder was the best-performing docking method in that challenge. BioMolTech are actively building on this excellent result with the aim of making robust and reliable activity predictions a standard outcome of a Lead Finder experiment.
Cresset are proud to be the worldwide distributors for Lead Finder. It is available today as a command-line application and will be built into Cresset’s upcoming structure-based drug design workbench.