The adaptability of pyFlare to access advanced data visualization and calculation functionalities
We showcase some examples within a drug discovery context where the pyFlare environment has been further expanded with advanced data ...
News
Cresset is very well known for providing fast and accurate ligand-based virtual screening through Blaze. We have now added the Lead Finder docking engine to our virtual screening toolbox, giving Cresset Discovery Services (CDS) the most comprehensive virtual screening capabilities available anywhere in the industry.
Based on an informal survey of our contacts and customers, I estimate that something like 50% of all current pharma SME projects are ‘structure enabled’. Lead discovery and lead optimization are driven through the use of in-house structures, public structures (typically from the PDB) and homology models. These structures inform lead optimization programs by explaining observed SAR and providing feedback and a detailed context for the design of further analogues.
CDS routinely uses the Cresset software Blaze for ligand-based virtual screening. Although we had access to structure-based methods, we are pleased to have brought Lead Finder in-house, giving us full capability in conducting ligand-protein docking.
Virtual screening with Blaze remains one of the most consistently requested projects for CDS. What makes Blaze extremely useful for our customers is:
Blaze also relies on two very simple premises:
So, the key determinants of real activity obtained from hit lists (other than was this truly the ‘bioactive conformation’?) is often just how relevant and what distribution that hit conformation has in the population. This is fundamentally why our ligand-centric screening invariably works extremely well. Given that a molecule can adopt a similar shape, and project the same electrostatic patterns, from a completely different chemical architecture, leads to a very diverse output.
The Lead Finder software has been developed to provide cutting-edge docking for an array of typical tasks, from high-throughput virtual screening to best-in-class prediction of bioactive conformations to accurate prediction of binding energies. In combination with the companion Build Model protein preparation tool, Lead Finder has been shown to match or outperform the historically leading docking solutions.
When preparing ligands for virtual screening in Blaze, CDS scientists use modeling to help define the best ‘hand-crafted’ estimate of a bioactive conformation, based on the widest data for any given system. We apply the same care to exploring and preparing protein targets prior to structure-based virtual screens. We take advantage of three main approaches. Firstly, Lead Finder includes the excellent Build Model protein preparation tool. Secondly, we are privileged to be able to model proteins and ligands using the same proprietary XED force field used to give the accurate electrostatics that all Cresset software is based on. Finally, at CDS we have access to the latest Cresset software that is still under development. This gives us capability to provide protein electrostatic field maps and water analysis, providing a very reliable starting position for structure-based virtual screening.
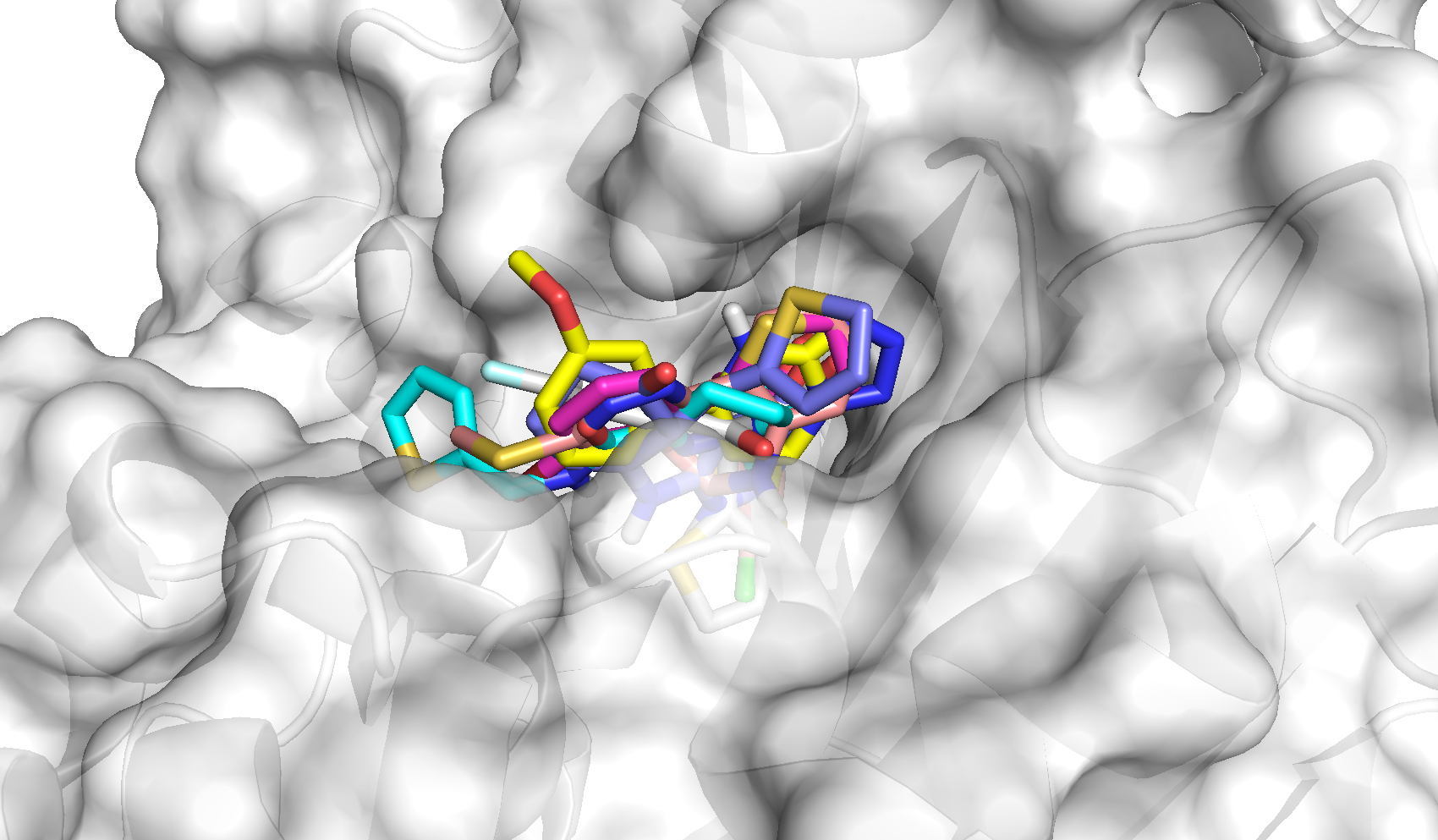
Lead Finder uses a stochastic ligand sampling workflow, with conformations generated on-the-fly, and a genetic algorithm for processing these into pools of the best docking poses. Multiple interaction grids are generated from the protein target and combined to define a scoring system for poses. More importantly, the scoring method has been shown to outperform some of the more conventional docking engines currently available commercially.
What are the advantages of having structure-based and ligand-based virtual screening? And how do we choose which is the best approach for a project?
Ligand-based virtual screening is less computationally intensive, making it a preferred option when there is a known ligand available. An average protein of 400 amino acids has over 20,000 heavy atoms and 9,600 bonds and in excess of 50 charges, making it a more challenging system to model.
However, even when there is a known ligand there are some situations when a ligand-based virtual screening is not viable, such as when the known ligand does not exploit all the interactions available in an active site or when a protein has an unattractive orthosteric site and attractive allosteric sites with no known ligands. In these cases, we prefer to use a structure-based method.
In the case of protein-protein interaction sites and protein-DNA/RNA sites, Blaze can take DNA and protein fragments as a template in place of a ligand. However, it is useful to have a structure-based approach available for comparison.
In fact, we often find it useful to combine different virtual screening techniques. In lead discovery, one of the key requirements for virtual screening is to maximise the diversity of hits returned. All virtual screening techniques, be they ligand-based or structure-based, are probabilistic techniques in that they may be used to increase the likelihood of getting hits from a wet screen. No technique guarantees to give absolute binding energies (at least not in the context of virtual screening on any realistic size of screening library), but they do give good rank ordering of compounds and can, therefore, be used as a means of selection and prioritisation.
Ligand-based techniques, whether 2D or 3D, are algorithmically distinct from structure-based techniques such as docking and, therefore, give different rankings to compounds. Different approaches return different hits and the results can be combined into an enriched final list.
Combining the results of structure-based and ligand-based techniques provides further diversity, leading to better hit rates and more interesting hits.
Through combining the strengths of Blaze in the ligand-based world with Lead Finder for docking, CDS now has the most comprehensive virtual screening capabilities available anywhere in the industry. Both Blaze and Lead Finder are available to purchase as software or as a service through CDS. CDS is truly now a one stop shop for virtual screening and indeed very much more.