Improving PROTAC properties via single-point changes to linkers
We explore how computational methods can be applied to proteolysis targeting chimera (PROTAC) design, to effectively tackle some of the ...
News
New versions (V10.5) of Forge™ and Torch™ are due out next month. This release offers new science and functionality and plenty of improvements that significantly enhance both applications. Below is a sneak peek at some of the new functionality in Forge.
In this release of Forge we have included the new options to constrain the alignments using specific pharmacophoric features. As in Blaze, constraints (e.g., DonorH, Acceptor, Cation, Anion, covalent center) can be added to reference molecules and must be matched in the alignment or a penalty will be applied to the score. Pharmacophore constraints will be useful in those cases (such as specific kinase targets or metal chelators) where explicit interactions dominate the alignments.
Molecule alignment is significantly improved in V10.5. New and enhanced functionality include:
The result of all these improvements will be significantly improved generation of alignments that match your expectations without manual interference.
Molecular conformations are central to what we do. We think that our conformation hunter does a good job of generating a diverse range of energetically accessible conformations. However, we wanted to give you the opportunity to more easily explore the conformations of your molecules, enabling you to interact with and edit the populations.
The conformation explorer is a new tool in Forge for visualizing and analyzing conformation analysis results. Within the conformation explorer you can:
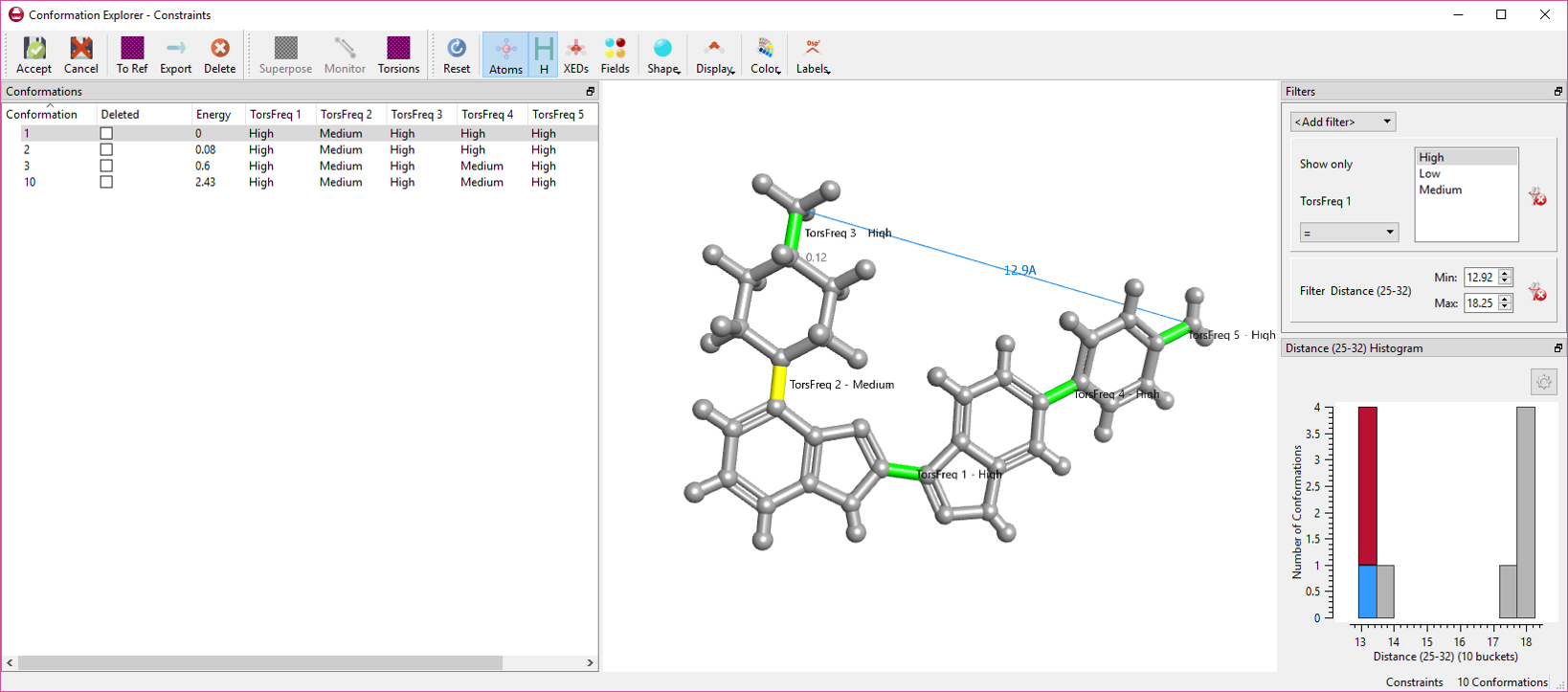
Figure 1: The conformation explorer in Forge. Rotatable bonds are colored and labelled by CSD torsion frequency.
Contact us to register for a free evaluation of Forge V10.5.