Improving PROTAC properties via single-point changes to linkers
We explore how computational methods can be applied to proteolysis targeting chimera (PROTAC) design, to effectively tackle some of the ...
News
In the releases of Blaze V10.3 and Forge V10.5 we introduced new similarity metrics alongside the new capabilities to manually weight the similarity function using pharmacophore constraints. With the introduction of Tanimoto and particularly Tversky measures of similarity, a new range of experiments are available to you that help you tailor the results you get. In this post I will use the Tversky similarity to perform substructure and superstructure type searches using Blaze. These new options are also available in Forge.
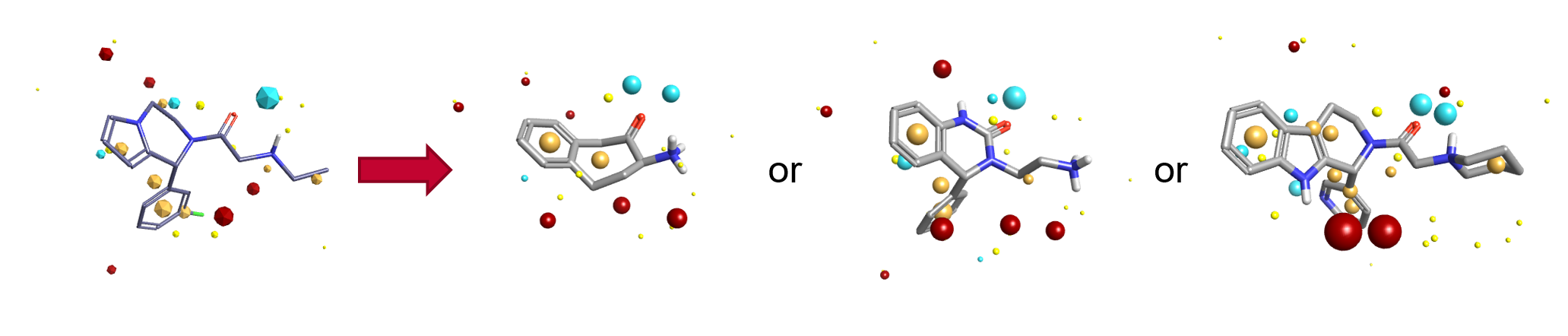
Figure 1: Blaze results can be tailored to generate the type of results that interest you, from substructure like to pure chemotype switching or super-structure like.
Blaze uses the field point patterns of molecules combined with their shape to align and score a ‘database’ of molecules against a ‘reference’ or ‘query’ that is usually a known active. In this context the default Dice similarity has worked well. It returns active molecules that are similar in size to the query, but is not too size-dependent allowing Blaze to find hits that are smaller than the reference. In most cases this is exactly what you want – a ligand the same size or smaller than the reference that maintains most of the potential sites of interaction. The scoring algorithm could be altered to generate more substructure like or more superstructure like results. However, this was complex to set up and sub-optimal in performance. In Blaze V10.3 the new Tversky similarity makes these searches more accessible. A look at the average MW of the first 100 compounds returned using the standard Dice and the new Tversky options highlights the difference:
Table of average MW of first 100 compounds returned using different similarity metrics. Database of 35283 positively charged Chembl compounds with 5-30 heavy atoms on Blaze. Query MW: 319. Database average MW: 318
| Dice | Tanimoto | Tversky, α 0.05 | Tversky, α 0.95 |
|---|---|---|---|
| 314 | 313 | 192 | 363 |
The Tversky metric has two parameters, α and β. Using the Tversky similarity option in Blaze, and setting α to 0.05 and β 0.95, results in a substructure-like search. In fact, we don’t deal with structures so this actually equates to a ‘sub-field’ search. It returns molecules that contain a field pattern that is contained within the query – i.e. field fragments of the query. This is useful where you have a large known active but want to screen or design a fragment library of smaller molecules that match parts of the query.
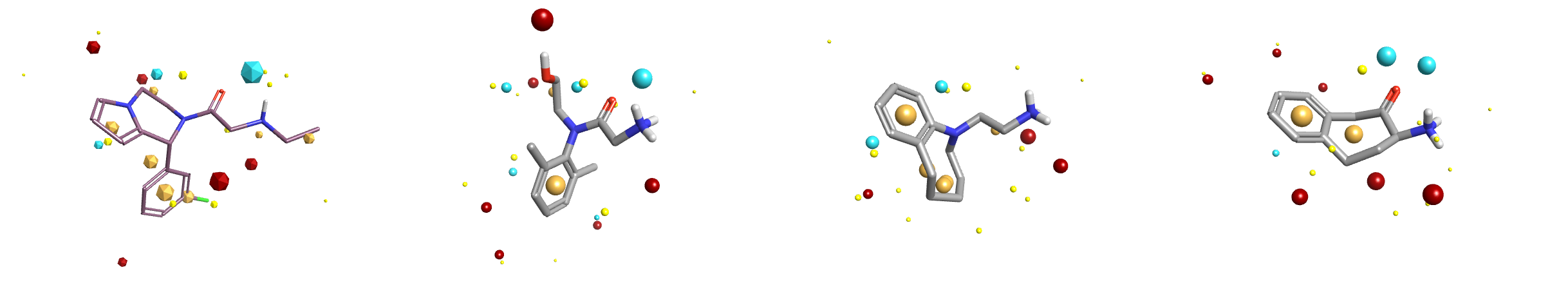
Figure 2: Search query and 3 selected results (ranks 3, 5, 11) from a sub-field search using the A2C active from the Fragment hopping with Blaze case study. Each result includes some features of the search query but also omits at least one functional group.
Setting a Tversky similarity with α at 0.95 and β at 0.05 generates a ‘super-field’ search. That is, molecules that contain a field pattern similar to the query are scored highly whether or not they have additional field points. This is useful for growing hits from a fragment screen or in other situations where you do not want to penalize results for having additional functionality to the query. As hits could contain the query at any position and any orientation, this option works particularly well when combined with field, pharmacophore or excluded volume constraints. For example, using an excluded volume will direct the results towards the available space around the query. Equally, using field constraints or the new pharmacophore constraints will ensure that results contain the interactions that you know to be important.
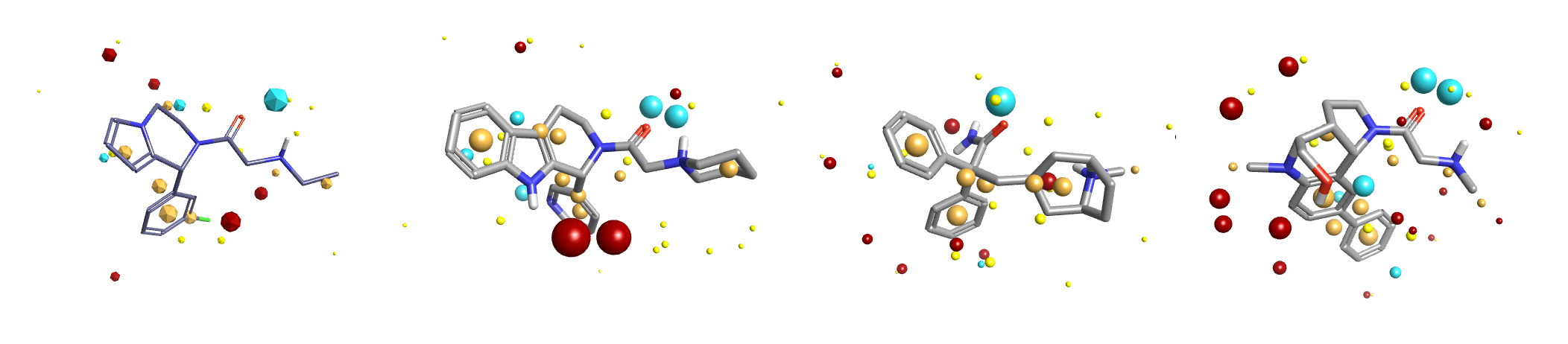
Figure 3: Search query and 3 selected results (ranks 2, 4, 6) from a super-field search using A2C active from the Fragment hopping with Blaze case study and an expanded database to include larger fragments. Each result contains a similar field pattern to the query plus additional features or functional groups.